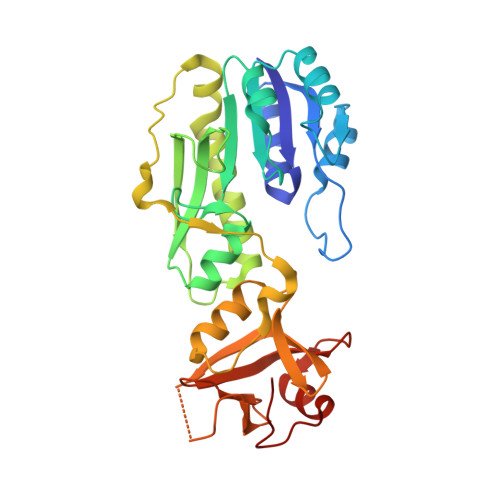
Structure of the Escherichia coli ArnA N-formyltransferase domain in complex with N(5) -formyltetrahydrofolate and UDP-Ara4N.
Genthe, N.A., Thoden, J.B., Holden, H.M.(2016) Protein Sci 25: 1555-1562
- PubMed: 27171345
- DOI: https://doi.org/10.1002/pro.2938
- Primary Citation of Related Structures:
5J63 - PubMed Abstract:
ArnA from Escherichia coli is a key enzyme involved in the formation of 4-amino-4-deoxy-l-arabinose. The addition of this sugar to the lipid A moiety of the lipopolysaccharide of pathogenic Gram-negative bacteria allows these organisms to evade the cationic antimicrobial peptides of the host immune system. Indeed, it is thought that such modifications may be responsible for the repeated infections of cystic fibrosis patients with Pseudomonas aeruginosa. ArnA is a bifunctional enzyme with the N- and C-terminal domains catalyzing formylation and oxidative decarboxylation reactions, respectively. The catalytically competent cofactor for the formylation reaction is N(10) -formyltetrahydrofolate. Here we describe the structure of the isolated N-terminal domain of ArnA in complex with its UDP-sugar substrate and N(5) -formyltetrahydrofolate. The model presented herein may prove valuable in the development of new antimicrobial therapeutics.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, Wisconsin, 53706.