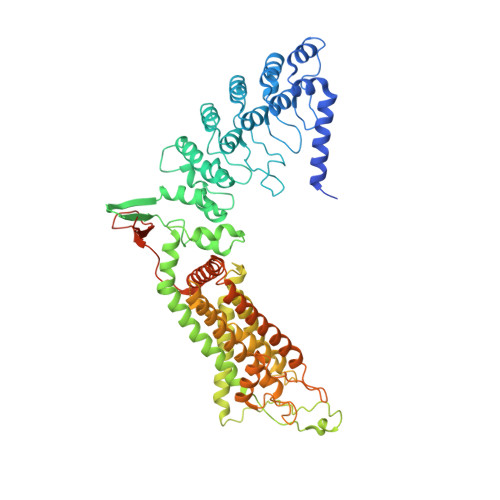
Crystal structure of the epithelial calcium channel TRPV6.
Saotome, K., Singh, A.K., Yelshanskaya, M.V., Sobolevsky, A.I.(2016) Nature 534: 506-511
- PubMed: 27296226
- DOI: https://doi.org/10.1038/nature17975
- Primary Citation of Related Structures:
5IWK, 5IWP, 5IWR, 5IWT - PubMed Abstract:
Precise regulation of calcium homeostasis is essential for many physiological functions. The Ca(2+)-selective transient receptor potential (TRP) channels TRPV5 and TRPV6 play vital roles in calcium homeostasis as Ca(2+) uptake channels in epithelial tissues. Detailed structural bases for their assembly and Ca(2+) permeation remain obscure. Here we report the crystal structure of rat TRPV6 at 3.25 Å resolution. The overall architecture of TRPV6 reveals shared and unique features compared with other TRP channels. Intracellular domains engage in extensive interactions to form an intracellular 'skirt' involved in allosteric modulation. In the K(+) channel-like transmembrane domain, Ca(2+) selectivity is determined by direct coordination of Ca(2+) by a ring of aspartate side chains in the selectivity filter. On the basis of crystallographically identified cation-binding sites at the pore axis and extracellular vestibule, we propose a Ca(2+) permeation mechanism. Our results provide a structural foundation for understanding the regulation of epithelial Ca(2+) uptake and its role in pathophysiology.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Columbia University, 650 West 168th Street, New York, New York 10032, USA.