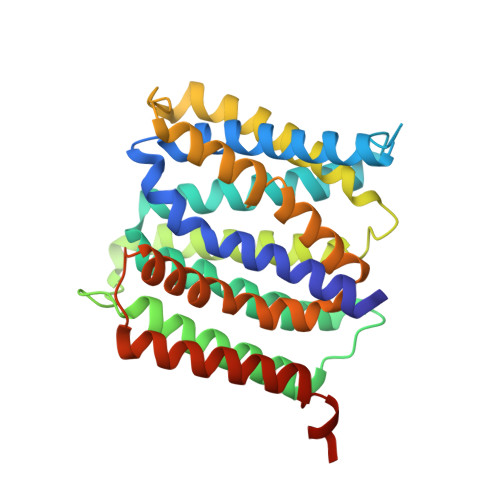
Structural basis for amino acid export by DMT superfamily transporter YddG.
Tsuchiya, H., Doki, S., Takemoto, M., Ikuta, T., Higuchi, T., Fukui, K., Usuda, Y., Tabuchi, E., Nagatoishi, S., Tsumoto, K., Nishizawa, T., Ito, K., Dohmae, N., Ishitani, R., Nureki, O.(2016) Nature 534: 417-420
- PubMed: 27281193
- DOI: https://doi.org/10.1038/nature17991
- Primary Citation of Related Structures:
5I20 - PubMed Abstract:
The drug/metabolite transporter (DMT) superfamily is a large group of membrane transporters ubiquitously found in eukaryotes, bacteria and archaea, and includes exporters for a remarkably wide range of substrates, such as toxic compounds and metabolites. YddG is a bacterial DMT protein that expels aromatic amino acids and exogenous toxic compounds, thereby contributing to cellular homeostasis. Here we present structural and functional analyses of YddG. Using liposome-based analyses, we show that Escherichia coli and Starkeya novella YddG export various amino acids. The crystal structure of S. novella YddG at 2.4 Å resolution reveals a new membrane transporter topology, with ten transmembrane segments in an outward-facing state. The overall structure is basket-shaped, with a large substrate-binding cavity at the centre of the molecule, and is composed of inverted structural repeats related by two-fold pseudo-symmetry. On the basis of this intramolecular symmetry, we propose a structural model for the inward-facing state and a mechanism of the conformational change for substrate transport, which we confirmed by biochemical analyses. These findings provide a structural basis for the mechanism of transport of DMT superfamily proteins.
Organizational Affiliation:
Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, Tokyo 113-0032, Japan.