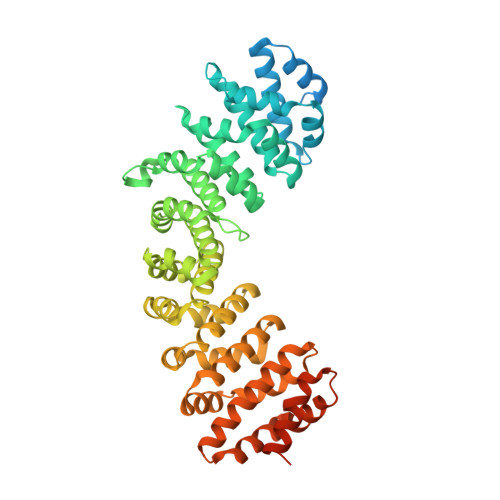
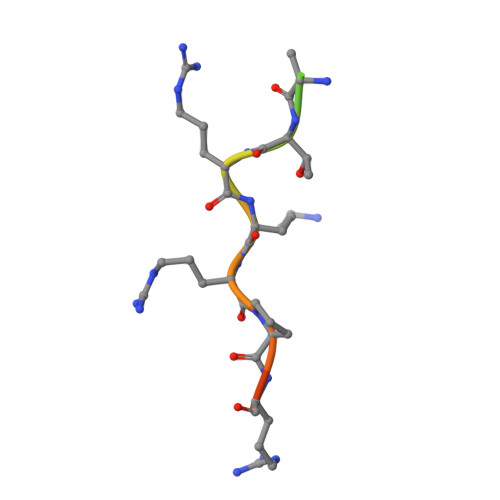
Divergent Evolution of Nuclear Localization Signal Sequences in Herpesvirus Terminase Subunits.
Sankhala, R.S., Lokareddy, R.K., Cingolani, G.(2016) J Biol Chem 291: 11420-11433
- PubMed: 27033706
- DOI: https://doi.org/10.1074/jbc.M116.724393
- Primary Citation of Related Structures:
5HUW, 5HUY - PubMed Abstract:
The tripartite terminase complex of herpesviruses assembles in the cytoplasm of infected cells and exploits the host nuclear import machinery to gain access to the nucleus, where capsid assembly and genome-packaging occur. Here we analyzed the structure and conservation of nuclear localization signal (NLS) sequences previously identified in herpes simplex virus 1 (HSV-1) large terminase and human cytomegalovirus (HCMV) small terminase. We found a monopartite NLS at the N terminus of large terminase, flanking the ATPase domain, that is conserved only in α-herpesviruses. In contrast, small terminase exposes a classical NLS at the far C terminus of its helical structure that is conserved only in two genera of the β-subfamily and absent in α- and γ-herpesviruses. In addition, we predicted a classical NLS in the third terminase subunit that is partially conserved among herpesviruses. Bioinformatic analysis revealed that both location and potency of NLSs in terminase subunits evolved more rapidly than the rest of the amino acid sequence despite the selective pressure to keep terminase gene products active and localized in the nucleus. We propose that swapping NLSs among terminase subunits is a regulatory mechanism that allows different herpesviruses to regulate the kinetics of terminase nuclear import, reflecting a mechanism of virus:host adaptation.
Organizational Affiliation:
From the Department of Biochemistry and Molecular Biology, Thomas Jefferson University, Philadelphia, Pennsylvania 19107 and.