Discovery of GPX4 inhibitory peptides from random peptide T7 phage display and subsequent structural analysis
Sakamoto, K., Sogabe, S., Kamada, Y., Matsumoto, S.I., Kadotani, A., Sakamoto, J.I., Tani, A.(2017) Biochem Biophys Res Commun 482: 195-201
- PubMed: 27836545
- DOI: https://doi.org/10.1016/j.bbrc.2016.11.035
- Primary Citation of Related Structures:
5H5Q, 5H5R, 5H5S - PubMed Abstract:
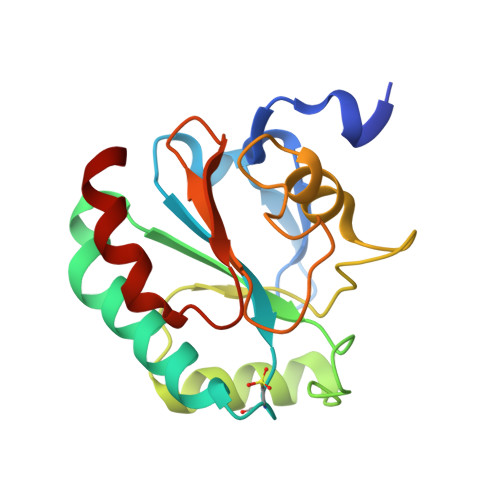
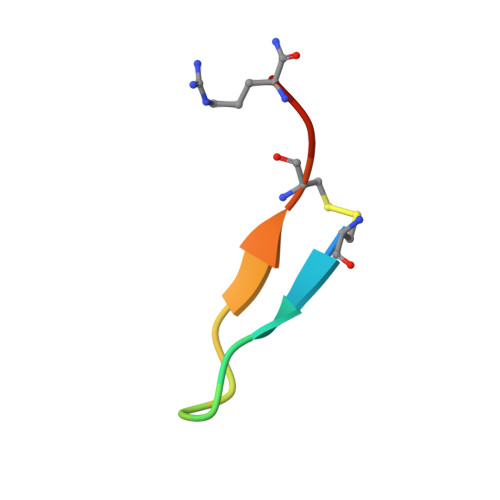
The phospholipid hydroperoxidase glutathione peroxidase (GPX4) is an enzyme that reduces lipid hydroperoxides in lipid membranes. Recently, GPX4 has been investigated as a target molecule that induces iron-dependent cell death (ferroptosis) selectively in cancer cells that express mutant Ras. GPX4 inhibitors have the potential to become novel anti-cancer drugs. However, there are no druggable pockets for conventional small molecules on the molecular surface of GPX4. To generate GPX4 inhibitors, we examined the use of peptides as an alternative to small molecules. By screening peptide libraries displayed on T7 phages, and analyzing the X-ray crystal structures of the peptides, we successfully identified one peptide that binds to near Sec73 of catalytic site and two peptides that bind to another site on GPX4. To our knowledge, this is the first study reporting GPX4 inhibitory peptides and their structural information.
Organizational Affiliation:
Pharmaceutical Research Division, Takeda Pharmaceutical Company Limited, 26-1, Muraoka-Higashi 2-chome, Fujisawa, Kanagawa, 251-8555, Japan. Electronic address: koutarou.sakamoto@takeda.com.