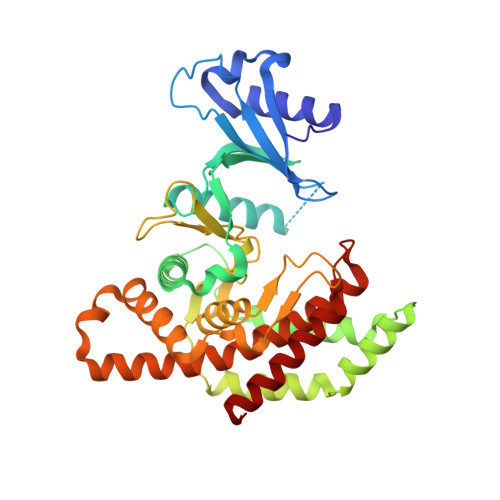
Design, Synthesis, Crystallization and Biological Evaluation of New Symmetrical Biscationic Compounds as Selective Inhibitors of Human Choline Kinase Alpha1 (Chokalpha1)
Schiaffino-Ortega, S., Baglioni, E., Mariotto, E., Bortolozzi, R., Serran-Aguilera, L., Rios-Marco, P., Carrasco-Jimenez, M.P., Gallo, M.A., Hurtado-Guerrero, R., Marco, C., Basso, G., Viola, G., Entrena, A., Lopez-Cara, L.C.(2016) Sci Rep 6: 23793
- PubMed: 27029499
- DOI: https://doi.org/10.1038/srep23793
- Primary Citation of Related Structures:
5FTG - PubMed Abstract:
A novel family of compounds derivative of 1,1'-(((ethane-1,2-diylbis(oxy))bis(4,1-phenylene))bis(methylene))-bispyridinium or -bisquinolinium bromide (10a-l) containing a pair of oxygen atoms in the spacer of the linker between the biscationic moieties, were synthesized and evaluated as inhibitors of choline kinase against a panel of cancer-cell lines. The most promising compounds in this series were 1,1'-(((ethane-1,2-diylbis(oxy))bis(4,1-phenylene))bis(methylene))bis(4-(dimethylamino)pyridinium) bromide (10a) and 1,1'-(((ethane-1,2-diylbis(oxy))bis(4,1-phenylene))bis(methylene))-bis(7-chloro-4-(pyrrolidin-1-yl)quinolinium) bromide (10l), which inhibit human choline kinase (ChoKα1) with IC50 of 1.0 and 0.92 μM, respectively, in a range similar to that of the previously reported biscationic compounds MN58b and RSM932A. Our compounds show greater antiproliferative activities than do the reference compounds, with unprecedented values of GI50 in the nanomolar range for several of the cancer-cell lines assayed, and more importantly they present low toxicity in non-tumoral cell lines, suggesting a cancer-cell-selective antiproliferative activity. Docking studies predict that the compounds interact with the choline-binding site in agreement with the binding mode of most previously reported biscationic compounds. Moreover, the crystal structure of ChoKα1 with compound 10a reveals that this compound binds to the choline-binding site and mimics HC-3 binding mode as never before.
Organizational Affiliation:
Departamento de Química Farmacéutica y Orgánica, Facultad de Farmacia, Campus de Cartuja, 18071 Granada (Spain).