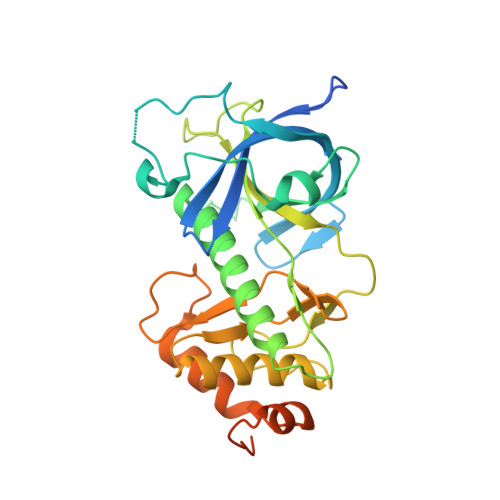
Structure of the Dictyostelium Myosin-II Heavy Chain Kinase A (MHCK-A) alpha-kinase domain apoenzyme reveals a novel autoinhibited conformation.
Ye, Q., Yang, Y., van Staalduinen, L., Crawley, S.W., Liu, L., Brennan, S., Cote, G.P., Jia, Z.(2016) Sci Rep 6: 26634-26634
- PubMed: 27211275
- DOI: https://doi.org/10.1038/srep26634
- Primary Citation of Related Structures:
5DYJ, 5E4H, 5E9E - PubMed Abstract:
The α-kinases are a family of a typical protein kinases present in organisms ranging from protozoa to mammals. Here we report an autoinhibited conformation for the α-kinase domain of Dictyostelium myosin-II heavy chain kinase A (MHCK-A) in which nucleotide binding to the catalytic cleft, located at the interface between an N-terminal and C-terminal lobe, is sterically blocked by the side chain of a conserved arginine residue (Arg592). Previous α-kinase structures have shown that an invariant catalytic aspartic acid residue (Asp766) is phosphorylated. Unexpectedly, in the autoinhibited conformation the phosphoryl group is transferred to the adjacent Asp663, creating an interaction network that stabilizes the autoinhibited state. The results suggest that Asp766 phosphorylation may play both catalytic and regulatory roles. The autoinhibited structure also provides the first view of a phosphothreonine residue docked into the phospho-specific allosteric binding site (Pi-pocket) in the C-lobe of the α-kinase domain.
Organizational Affiliation:
Department of Biomedical and Molecular Sciences, Queen's University, Kingston, ON, K7L 3N6, Canada.