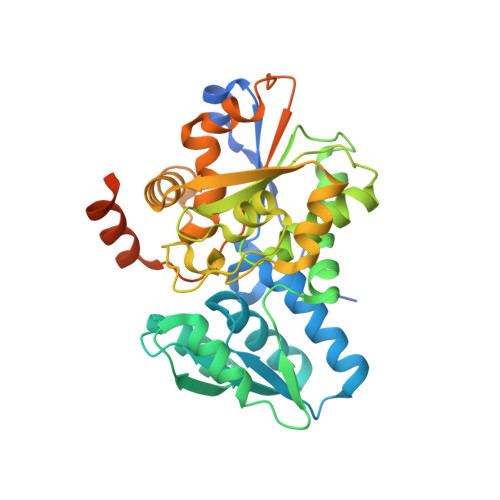
Crystal structure and characterization of a novel l-serine ammonia-lyase from Rhizomucor miehei.
Qin, Z., Yan, Q., Ma, Q., Jiang, Z.(2015) Biochem Biophys Res Commun 466: 431-437
- PubMed: 26367174
- DOI: https://doi.org/10.1016/j.bbrc.2015.09.043
- Primary Citation of Related Structures:
5C3U - PubMed Abstract:
L-serine ammonia-lyase, as a member of the β-family of pyridoxal-5'-phosphate (PLP) dependent enzymes, catalyzes the conversion of L-serine (L-threonine) to pyruvate (α-ketobutyrate) and ammonia. The crystal structure of L-serine ammonia-lyase from Rhizomucor miehei (RmSDH) was solved at 1.76 Å resolution by X-ray diffraction method. The overall structure of RmSDH had the characteristic β-family PLP dependent enzyme fold. It consisted of two distinct domains, both of which show the typical open twisted α/β structure. A PLP cofactor was located in the crevice between the two domains, which was attached to Lys52 by a Schiff-base linkage. Unique residue substitutions (Gly78, Pro79, Ser146, Ser147 and Thr312) were discovered at the catalytic site of RmSDH by comparison of structures of RmSDH and other reported eukaryotic L-serine ammonia-lyases. Optimal pH and temperature of the purified RmSDH were 7.5 and 40 °C, respectively. It was stable in the pH range of 7.0-9.0 and at temperatures below 40 °C. This is the first crystal structure of a fungal L-serine ammonia-lyase. It will be useful to study the catalytic mechanism of β-elimination enzymes and will provide a basis for further enzyme engineering.
Organizational Affiliation:
College of Food Science and Nutritional Engineering, Beijing Advanced Innovation Center of Food Nutrition and Human Health, China Agricultural University, Beijing 100083, China.