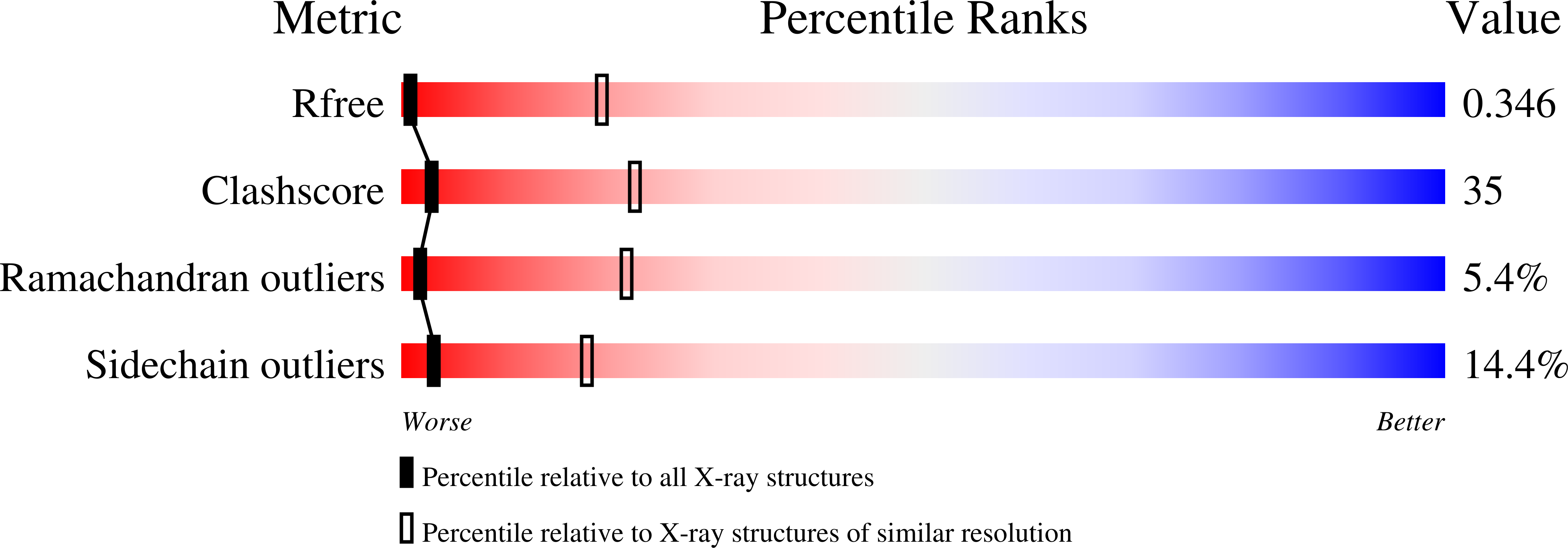
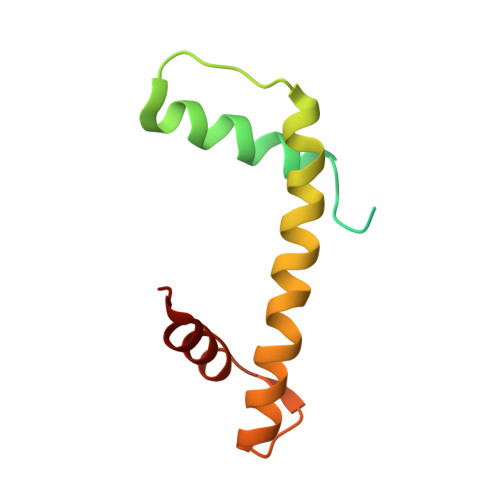
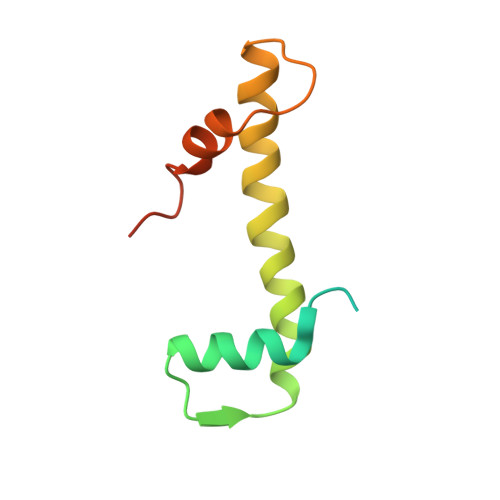
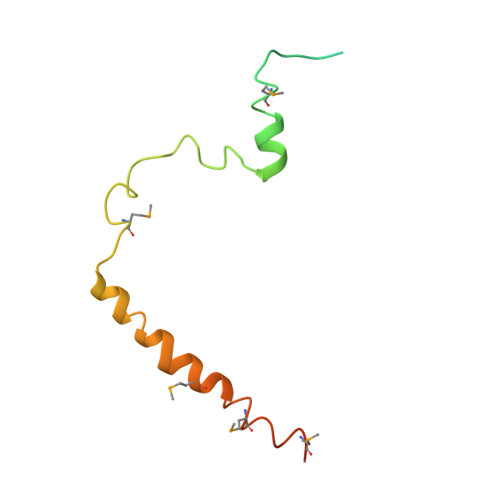
Structure-function studies of histone H3/H4 tetramer maintenance during transcription by chaperone Spt2.
Chen, S., Rufiange, A., Huang, H., Rajashankar, K.R., Nourani, A., Patel, D.J.(2015) Genes Dev 29: 1326-1340
- PubMed: 26109053
- DOI: https://doi.org/10.1101/gad.261115.115
- Primary Citation of Related Structures:
5BS7, 5BSA - PubMed Abstract:
Cells use specific mechanisms such as histone chaperones to abrogate the inherent barrier that the nucleosome poses to transcribing polymerases. The current model postulates that nucleosomes can be transiently disrupted to accommodate passage of RNA polymerases and that histones H3 and H4 possess their own chaperones dedicated to the recovery of nucleosomes. Here, we determined the crystal structure of the conserved C terminus of human Suppressors of Ty insertions 2 (hSpt2C) chaperone bound to an H3/H4 tetramer. The structural studies demonstrate that hSpt2C is bound to the periphery of the H3/H4 tetramer, mimicking the trajectory of nucleosomal-bound DNA. These structural studies have been complemented with in vitro binding and in vivo functional studies on mutants that disrupt key intermolecular contacts involving two acidic patches and hydrophobic residues on Spt2C. We show that contacts between both human and yeast Spt2C with the H3/H4 tetramer are required for the suppression of H3/H4 exchange as measured by H3K56ac and new H3 deposition. These interactions are also crucial for the inhibition of spurious transcription from within coding regions. Together, our data indicate that Spt2 interacts with the periphery of the H3/H4 tetramer and promotes its recycling in the wake of RNA polymerase.
Organizational Affiliation:
Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, New York 10065, USA;