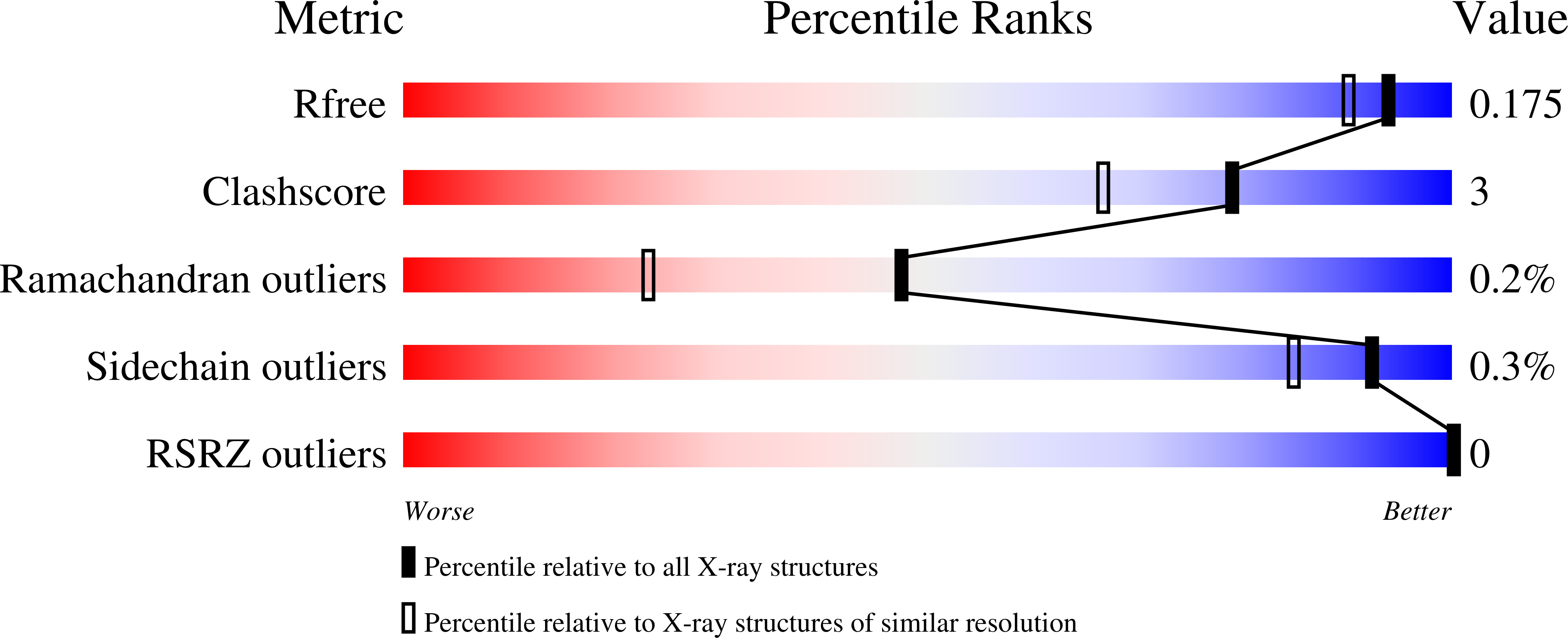
1.55 Angstrom Resolution Crystal Structure of Glutathione Reductase from Yersinia pestis in Complex with FAD.
Minasov, G., Shuvalova, L., Dubrovska, I., Cardona-Correa, A., Grimshaw, S., Kwon, K., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.