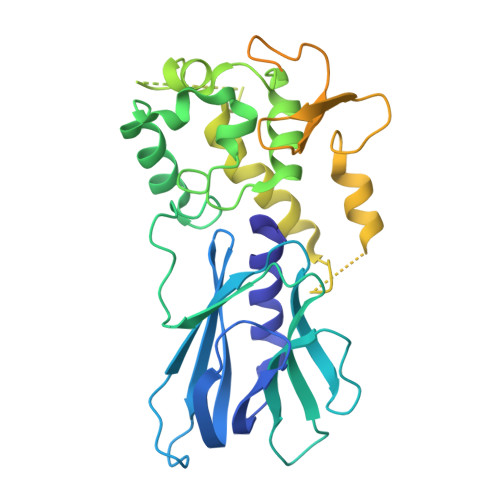
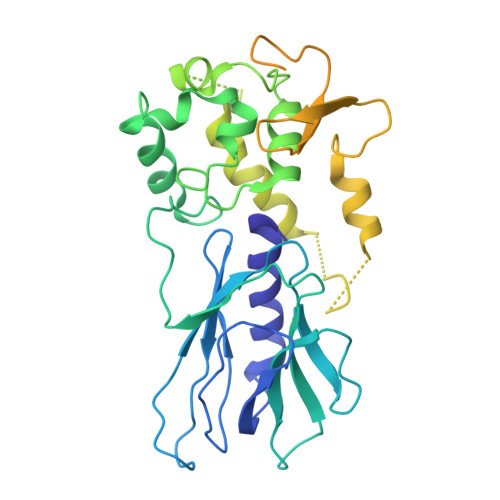
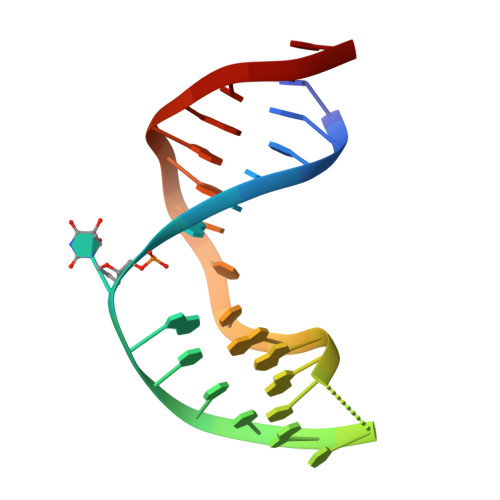
Tautomerization-dependent recognition and excision of oxidation damage in base-excision DNA repair
Zhu, C., Lu, L., Zhang, J., Yue, Z., Song, J., Zong, S., Liu, M., Stovicek, O., Gao, Y.Q., Yi, C.(2016) Proc Natl Acad Sci U S A 113: 7792-7797
- PubMed: 27354518
- DOI: https://doi.org/10.1073/pnas.1604591113
- Primary Citation of Related Structures:
5ITQ, 5ITR, 5ITT, 5ITU, 5ITX, 5ITY - PubMed Abstract:
NEIL1 (Nei-like 1) is a DNA repair glycosylase guarding the mammalian genome against oxidized DNA bases. As the first enzymes in the base-excision repair pathway, glycosylases must recognize the cognate substrates and catalyze their excision. Here we present crystal structures of human NEIL1 bound to a range of duplex DNA. Together with computational and biochemical analyses, our results suggest that NEIL1 promotes tautomerization of thymine glycol (Tg)-a preferred substrate-for optimal binding in its active site. Moreover, this tautomerization event also facilitates NEIL1-catalyzed Tg excision. To our knowledge, the present example represents the first documented case of enzyme-promoted tautomerization for efficient substrate recognition and catalysis in an enzyme-catalyzed reaction.
Organizational Affiliation:
State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing 100871, China;