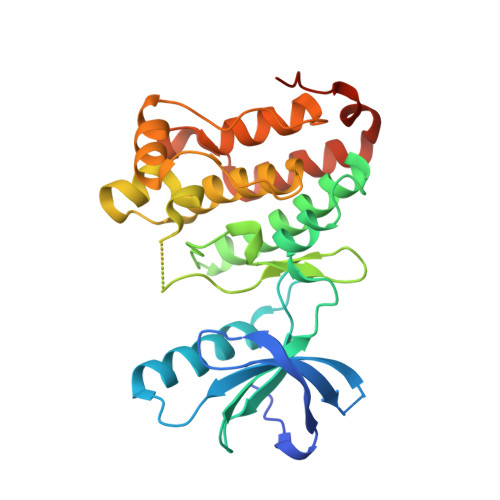
Chemical Proteomics and Structural Biology Define EPHA2 Inhibition by Clinical Kinase Drugs.
Heinzlmeir, S., Kudlinzki, D., Sreeramulu, S., Klaeger, S., Gande, S.L., Linhard, V., Wilhelm, M., Qiao, H., Helm, D., Ruprecht, B., Saxena, K., Medard, G., Schwalbe, H., Kuster, B.(2016) ACS Chem Biol 11: 3400-3411
- PubMed: 27768280
- DOI: https://doi.org/10.1021/acschembio.6b00709
- Primary Citation of Related Structures:
5I9U, 5I9V, 5I9W, 5I9X, 5I9Y, 5I9Z, 5IA0, 5IA1, 5IA2, 5IA3, 5IA4, 5IA5 - PubMed Abstract:
The receptor tyrosine kinase EPHA2 (Ephrin type-A receptor 2) plays important roles in oncogenesis, metastasis, and treatment resistance, yet therapeutic targeting, drug discovery, or investigation of EPHA2 biology is hampered by the lack of appropriate inhibitors and structural information. Here, we used chemical proteomics to survey 235 clinical kinase inhibitors for their kinase selectivity and identified 24 drugs with submicromolar affinities for EPHA2. NMR-based conformational dynamics together with nine new cocrystal structures delineated drug-EPHA2 interactions in full detail. The combination of selectivity profiling, structure determination, and kinome wide sequence alignment allowed the development of a classification system in which amino acids in the drug binding site of EPHA2 are categorized into key, scaffold, potency, and selectivity residues. This scheme should be generally applicable in kinase drug discovery, and we anticipate that the provided information will greatly facilitate the development of selective EPHA2 inhibitors in particular and the repurposing of clinical kinase inhibitors in general.
Organizational Affiliation:
Chair of Proteomics and Bioanalytics, Technical University of Munich , 85354 Freising, Germany.