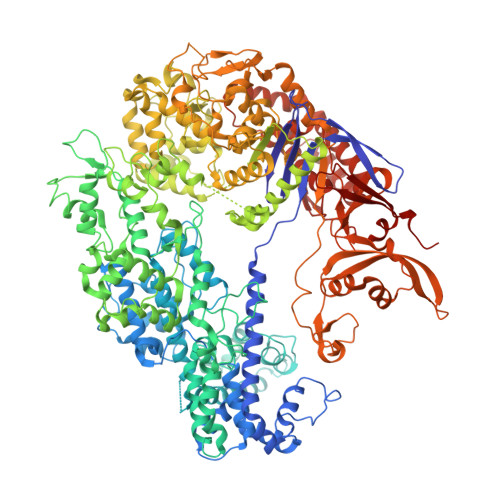
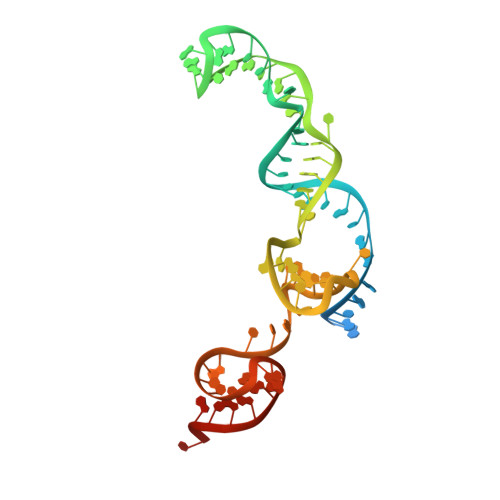
A Cas9-guide RNA complex preorganized for target DNA recognition.
Jiang, F., Zhou, K., Ma, L., Gressel, S., Doudna, J.A.(2015) Science 348: 1477-1481
- PubMed: 26113724
- DOI: https://doi.org/10.1126/science.aab1452
- Primary Citation of Related Structures:
4ZT0, 4ZT9 - PubMed Abstract:
Bacterial adaptive immunity uses CRISPR (clustered regularly interspaced short palindromic repeats)-associated (Cas) proteins together with CRISPR transcripts for foreign DNA degradation. In type II CRISPR-Cas systems, activation of Cas9 endonuclease for DNA recognition upon guide RNA binding occurs by an unknown mechanism. Crystal structures of Cas9 bound to single-guide RNA reveal a conformation distinct from both the apo and DNA-bound states, in which the 10-nucleotide RNA "seed" sequence required for initial DNA interrogation is preordered in an A-form conformation. This segment of the guide RNA is essential for Cas9 to form a DNA recognition-competent structure that is poised to engage double-stranded DNA target sequences. We construe this as convergent evolution of a "seed" mechanism reminiscent of that used by Argonaute proteins during RNA interference in eukaryotes.
Organizational Affiliation:
Department of Molecular and Cell Biology, University of California, Berkeley, CA 94720, USA.