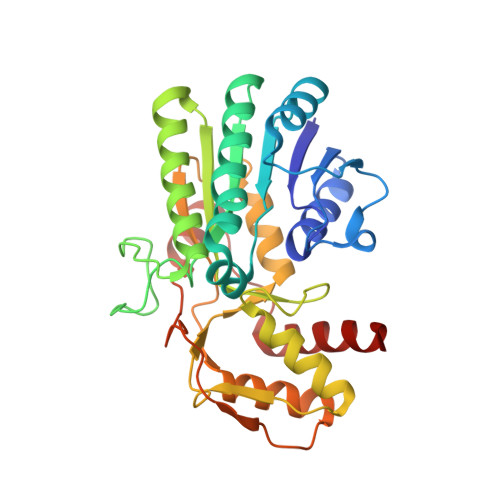
The structural basis of substrate promiscuity in UDP-hexose 4-epimerase from the hyperthermophilic Eubacterium Thermotoga maritima.
Shin, S.M., Choi, J.M., di Luccio, E., Lee, Y.J., Lee, S.J., Lee, S.J., Lee, S.H., Lee, D.W.(2015) Arch Biochem Biophys 585: 39-51
- PubMed: 26344854
- DOI: https://doi.org/10.1016/j.abb.2015.08.025
- Primary Citation of Related Structures:
4ZRM, 4ZRN - PubMed Abstract:
UDP-galactose 4-epimerase (GalE) catalyzes the interconversion of UDP-glucose (UDP-Glc) and UDP-galactose (UDP-Gal), which is a pivotal step in the Leloir pathway for d-galactose metabolism. Although GalE is widely distributed in prokaryotes and eukaryotes, little information is available regarding hyperthermophilic GalE. We overexpressed the TM0509 gene, encoding a putative GalE from Thermotoga maritima (TMGalE), in Escherichia coli and characterized the encoded protein. To further investigate the molecular basis of this enzyme's catalytic function, we determined the crystal structures of TMGalE and TMGalE bound to UDP-Glc at resolutions of 1.9 Å and 2.0 Å, respectively. The enzyme was determined to be a homodimer with a molecular mass of 70 kDa. The enzyme could reversibly catalyze the epimerization of UDP-GalNAc/UDP-GlcNAc as well as UDP-Gal/UDP-Glc at elevated temperatures, with an apparent optimal temperature and pH of 80 °C and 7.0, respectively. Our data showed that TM0509 is a UDP-galactosugar 4-epimerase involved in d-galactose metabolism; consequently, this study provides the first detailed characterization of a hyperthermophilic GalE. Moreover, the promiscuous substrate specificity of TMGalE, which is more similar to human GalE than E. coli GalE, supports the notion that TMGalE might exhibit the earliest form of sugar-epimerizing enzymes in the evolution of galactose metabolism.
Organizational Affiliation:
School of Applied Biosciences, Kyungpook National University, Daegu 702-701, South Korea.