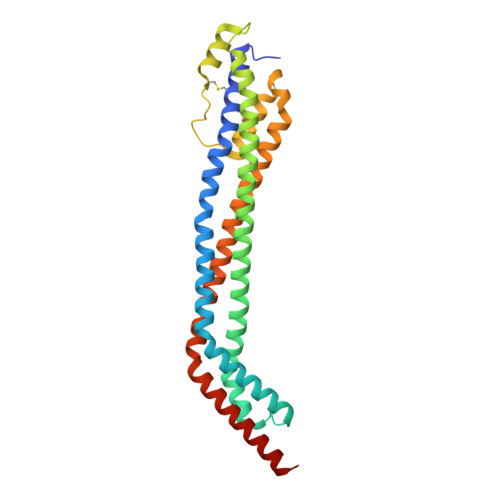
Structural basis for ligand and innate immunity factor uptake by the trypanosome haptoglobin-haemoglobin receptor.
Lane-Serff, H., MacGregor, P., Lowe, E.D., Carrington, M., Higgins, M.K.(2014) Elife 3: e05553-e05553
- PubMed: 25497229
- DOI: https://doi.org/10.7554/eLife.05553
- Primary Citation of Related Structures:
4X0J, 4X0L, 5HU6 - PubMed Abstract:
The haptoglobin-haemoglobin receptor (HpHbR) of African trypanosomes allows acquisition of haem and provides an uptake route for trypanolytic factor-1, a mediator of innate immunity against trypanosome infection. In this study, we report the structure of Trypanosoma brucei HpHbR in complex with human haptoglobin-haemoglobin (HpHb), revealing an elongated ligand-binding site that extends along its membrane distal half. This contacts haptoglobin and the β-subunit of haemoglobin, showing how the receptor selectively binds HpHb over individual components. Lateral mobility of the glycosylphosphatidylinositol-anchored HpHbR, and a ∼50° kink in the receptor, allows two receptors to simultaneously bind one HpHb dimer. Indeed, trypanosomes take up dimeric HpHb at significantly lower concentrations than monomeric HpHb, due to increased ligand avidity that comes from bivalent binding. The structure therefore reveals the molecular basis for ligand and innate immunity factor uptake by trypanosomes and identifies adaptations that allow efficient ligand uptake in the context of the complex trypanosome cell surface.
Organizational Affiliation:
Department of Biochemistry, University of Oxford, Oxford, United Kingdom.