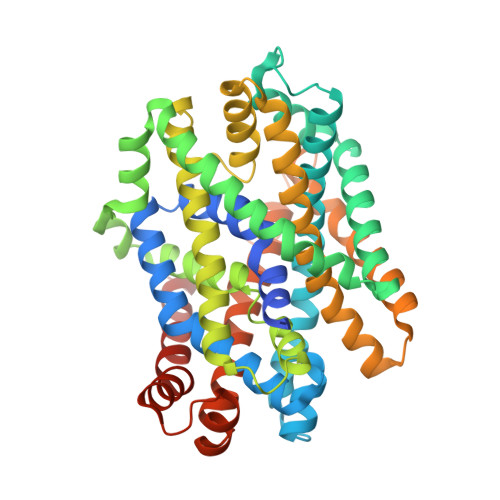
A Mechanism for Intracellular Release of Na+ by Neurotransmitter/Sodium Symporters
Malinauskaite, L., Quick, M., Reinhard, L., Lyons, J.A., Yano, H., Javitch, J.A., Nissen, P.(2014) Nat Struct Mol Biol 21: 1006
- PubMed: 25282149
- DOI: https://doi.org/10.1038/nsmb.2894
- Primary Citation of Related Structures:
4US3, 4US4 - PubMed Abstract:
Neurotransmitter/sodium symporters (NSSs) terminate synaptic signal transmission by Na+-dependent reuptake of released neurotransmitters. Key conformational states have been reported for the bacterial homolog LeuT and an inhibitor-bound Drosophila dopamine transporter. However, a coherent mechanism of Na+-driven transport has not been described. Here, we present two crystal structures of MhsT, an NSS member from Bacillus halodurans, in occluded inward-facing states with bound Na+ ions and L-tryptophan, providing insight into the cytoplasmic release of Na+. The switch from outward- to inward-oriented states is centered on the partial unwinding of transmembrane helix 5, facilitated by a conserved GlyX9Pro motif that opens an intracellular pathway for water to access the Na2 site. We propose a mechanism, based on our structural and functional findings, in which solvation through the TM5 pathway facilitates Na+ release from Na2 and the transition to an inward-open state.
Organizational Affiliation:
1] Danish Research Institute of Translational Neuroscience (DANDRITE), Nordic EMBL Partnership for Molecular Medicine, Aarhus University, Aarhus, Denmark. [2] Department of Molecular Biology and Genetics, Aarhus University, Aarhus, Denmark.