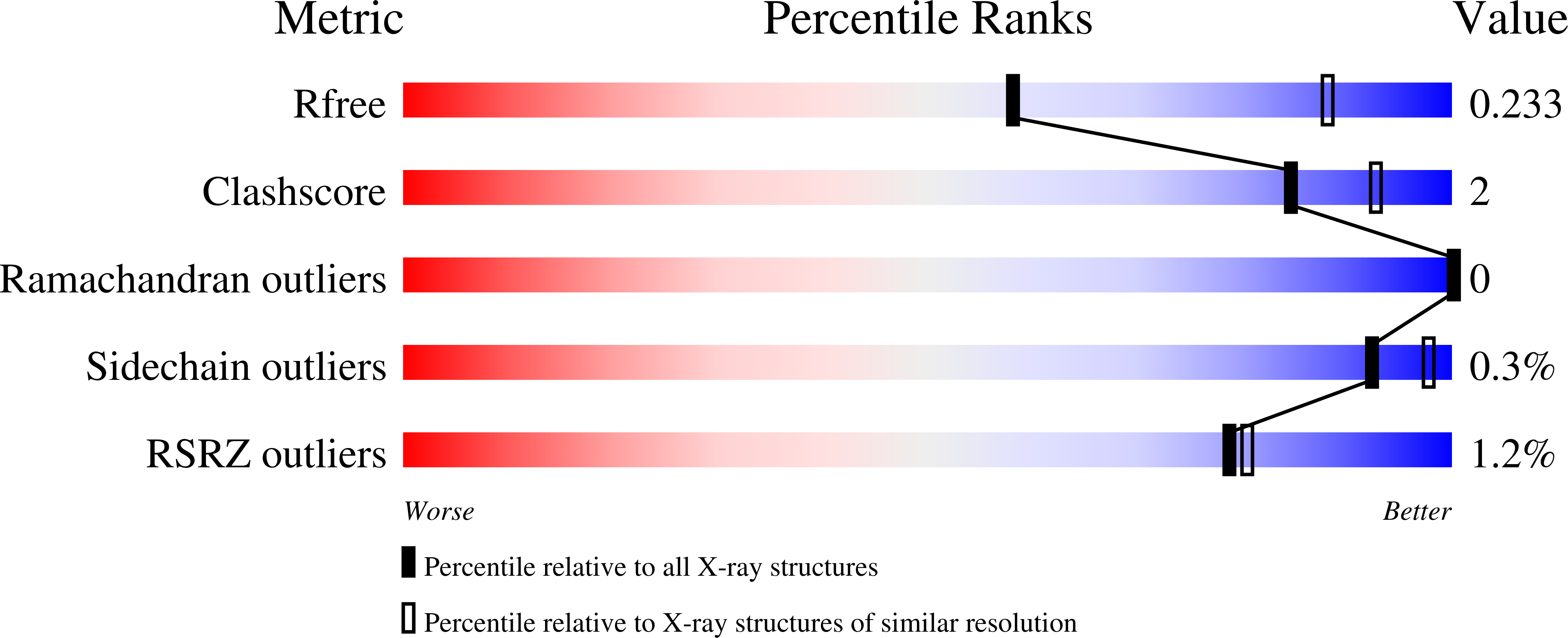
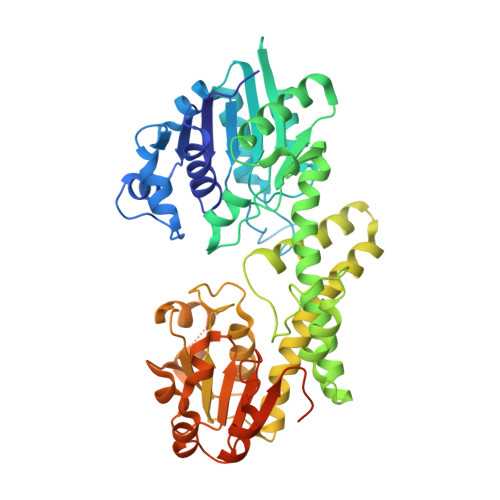
Hysteresis in Human UDP-Glucose Dehydrogenase Is Due to a Restrained Hexameric Structure That Favors Feedback Inhibition.
Kadirvelraj, R., Custer, G.S., Keul, N.D., Sennett, N.C., Sidlo, A.M., Walsh Jr., R.M., Wood, Z.A.(2014) Biochemistry 53: 8043-8051
- PubMed: 25478983
- DOI: https://doi.org/10.1021/bi500594x
- Primary Citation of Related Structures:
4RJT - PubMed Abstract:
Human UDP-α-d-glucose-6-dehydrogenase (hUGDH) displays hysteresis because of a slow isomerization from an inactive state (E*) to an active state (E). Here we show that the structure of E* constrains hUGDH in a conformation that favors feedback inhibition at physiological pH. The feedback inhibitor UDP-α-d-xylose (UDP-Xyl) competes with the substrate UDP-α-d-glucose for the active site. Upon binding, UDP-Xyl triggers an allosteric switch that changes the structure and affinity of the intersubunit interface to form a stable but inactive horseshoe-shaped hexamer. Using sedimentation velocity studies and a new crystal structure, we show that E* represents a stable conformational intermediate between the active and feedback-inhibited conformations. Because the allosteric switch occludes the cofactor and substrate binding sites in the inactive hexamer, the intermediate conformation observed in the crystal structure is consistent with the E* transient observed in relaxation studies. Steady-state analysis shows that the E* conformation enhances the affinity of hUGDH for the allosteric inhibitor UDP-Xyl by 8.6-fold (Ki = 810 nM). We present a model in which the constrained quaternary structure permits a small effector molecule to leverage a disproportionately large allosteric response.
Organizational Affiliation:
Department of Biochemistry & Molecular Biology, University of Georgia , Athens, Georgia 30602, United States.