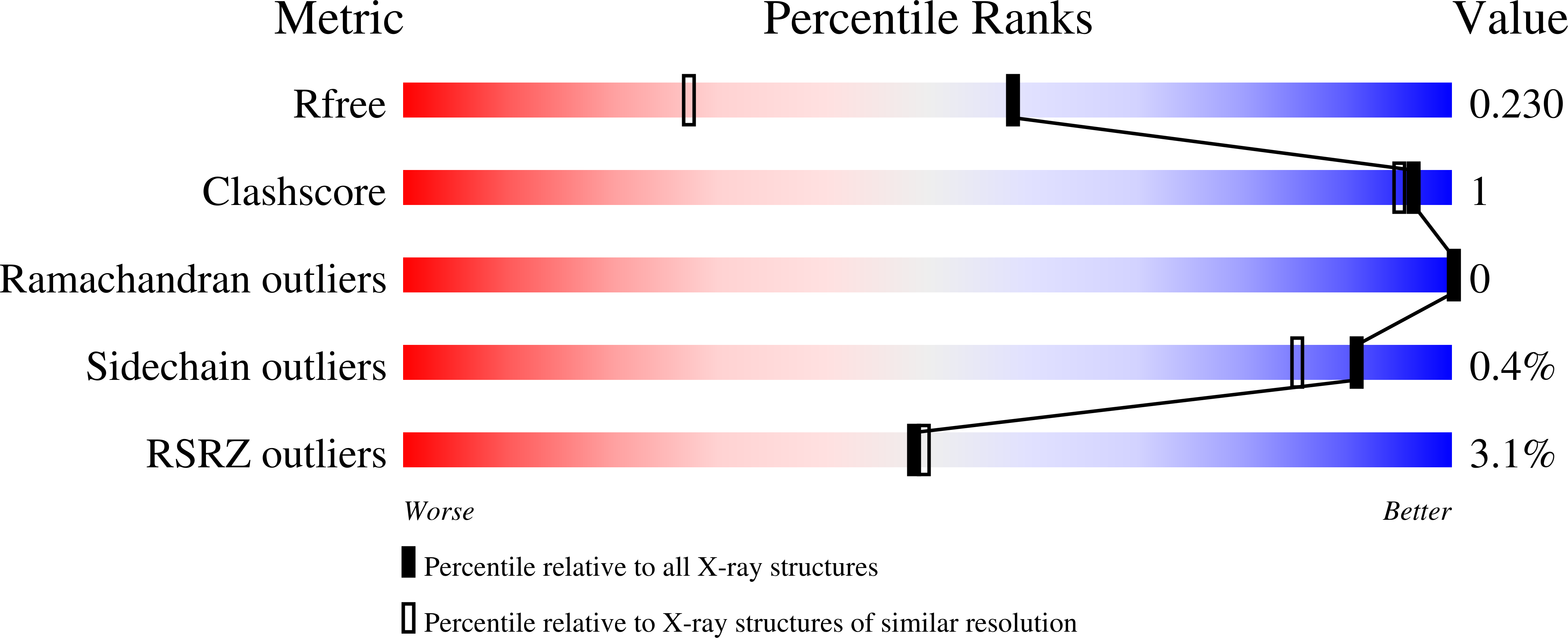
Phosphoglucan-bound structure of starch phosphatase Starch Excess4 reveals the mechanism for C6 specificity.
Meekins, D.A., Raththagala, M., Husodo, S., White, C.J., Guo, H.F., Kotting, O., Vander Kooi, C.W., Gentry, M.S.(2014) Proc Natl Acad Sci U S A 111: 7272-7277
- PubMed: 24799671
- DOI: https://doi.org/10.1073/pnas.1400757111
- Primary Citation of Related Structures:
4PYH - PubMed Abstract:
Plants use the insoluble polyglucan starch as their primary glucose storage molecule. Reversible phosphorylation, at the C6 and C3 positions of glucose moieties, is the only known natural modification of starch and is the key regulatory mechanism controlling its diurnal breakdown in plant leaves. The glucan phosphatase Starch Excess4 (SEX4) is a position-specific starch phosphatase that is essential for reversible starch phosphorylation; its absence leads to a dramatic accumulation of starch in Arabidopsis, but the basis for its function is unknown. Here we describe the crystal structure of SEX4 bound to maltoheptaose and phosphate to a resolution of 1.65 Å. SEX4 binds maltoheptaose via a continuous binding pocket and active site that spans both the carbohydrate-binding module (CBM) and the dual-specificity phosphatase (DSP) domain. This extended interface is composed of aromatic and hydrophilic residues that form a specific glucan-interacting platform. SEX4 contains a uniquely adapted DSP active site that accommodates a glucan polymer and is responsible for positioning maltoheptaose in a C6-specific orientation. We identified two DSP domain residues that are responsible for SEX4 site-specific activity and, using these insights, we engineered a SEX4 double mutant that completely reversed specificity from the C6 to the C3 position. Our data demonstrate that the two domains act in consort, with the CBM primarily responsible for engaging glucan chains, whereas the DSP integrates them in the catalytic site for position-specific dephosphorylation. These data provide important insights into the structural basis of glucan phosphatase site-specific activity and open new avenues for their biotechnological utilization.
Organizational Affiliation:
Department of Molecular and Cellular Biochemistry and Center for Structural Biology, University of Kentucky, Lexington, KY 40535-0509; and.