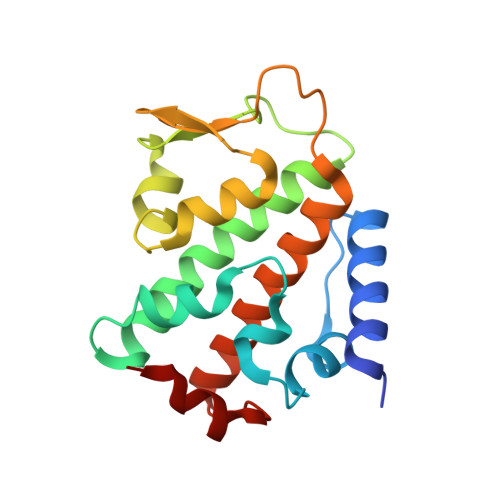
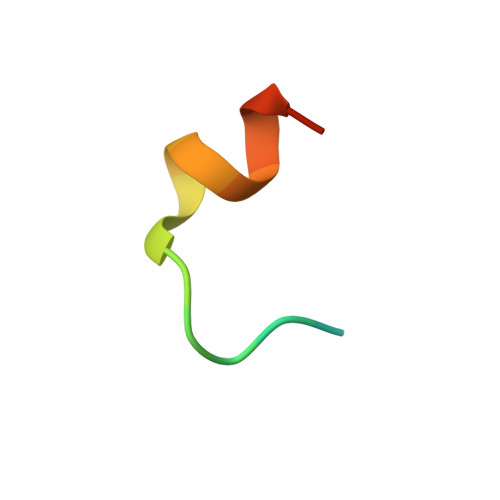
Molecular Basis and Regulation of OTULIN-LUBAC Interaction.
Elliott, P.R., Nielsen, S.V., Marco-Casanova, P., Fiil, B.K., Keusekotten, K., Mailand, N., Freund, S.M., Gyrd-Hansen, M., Komander, D.(2014) Mol Cell 54: 335-348
- PubMed: 24726323
- DOI: https://doi.org/10.1016/j.molcel.2014.03.018
- Primary Citation of Related Structures:
4OYJ, 4OYK - PubMed Abstract:
The linear ubiquitin (Ub) chain assembly complex (LUBAC) generates Met1-linked "linear" Ub chains that regulate the activation of the nuclear factor κB (NFκB) transcription factor and other processes. We recently discovered OTULIN as a deubiquitinase that specifically cleaves Met1-linked polyUb. Now, we show that OTULIN binds via a conserved PUB-interacting motif (PIM) to the PUB domain of the LUBAC component HOIP. Crystal structures and nuclear magnetic resonance experiments reveal the molecular basis for the high-affinity interaction and explain why OTULIN binds the HOIP PUB domain specifically. Analysis of LUBAC-induced NFκB signaling suggests that OTULIN needs to be present on LUBAC in order to restrict Met1-polyUb signaling. Moreover, LUBAC-OTULIN complex formation is regulated by OTULIN phosphorylation in the PIM. Phosphorylation of OTULIN prevents HOIP binding, whereas unphosphorylated OTULIN is part of the endogenous LUBAC complex. Our work exemplifies how coordination of ubiquitin assembly and disassembly activities in protein complexes regulates individual Ub linkage types.
Organizational Affiliation:
Medical Research Council Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge, CB2 0QH, UK.