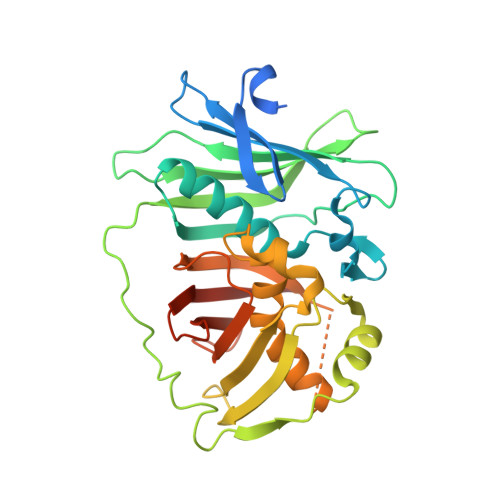
Structure and stereospecificity of the dehydratase domain from the terminal module of the rifamycin polyketide synthase.
Gay, D., You, Y.O., Keatinge-Clay, A., Cane, D.E.(2013) Biochemistry 52: 8916-8928
- PubMed: 24274103
- DOI: https://doi.org/10.1021/bi400988t
- Primary Citation of Related Structures:
4LN9 - PubMed Abstract:
RifDH10, the dehydratase domain from the terminal module of the rifamycin polyketide synthase, catalyzes the stereospecific syn dehydration of the model substrate (2S,3S)-2-methyl-3-hydroxypentanoyl-RifACP10, resulting in the exclusive formation of (E)-2-methyl-2-pentenoyl-RifACP10. RifDH10 does not dehydrate any of the other three diastereomeric, RifACP10-bound, diketide thioester substrates. On the other hand, when EryACP6, from the sixth module of the erythromycin polyketide synthase, is substituted for RifACP10, RifDH10 stereospecifically dehydrates only (2R,3R)-2-methyl-3-hydroxypentanoyl-EryACP6 to give exclusively (E)-2-methyl-2-pentenoyl-EryACP6, with no detectable dehydration of any of the other three diastereomeric, EryACP6-bound, diketides. An identical alteration in substrate diastereospecificity was observed for the corresponding N-acetylcysteamine or pantetheine thioester analogues, regardless of acyl chain length or substitution pattern. Incubation of (2RS)-2-methyl-3-ketopentanoyl-RifACP10 with the didomain reductase-dehydratase RifKR10-RifDH10 yielded (E)-2-methyl-2-pentenoyl-RifACP10, the expected product of syn dehydration of (2S,3S)-2-methyl-3-hydroxypentanoyl-RifACP10, while incubation with the corresponding EryACP6-bound substrate, (2RS)-2-methyl-3-ketopentanoyl-EryACP6, gave only the reduction product (2S,3S)-2-methyl-3-hydroxypentanoyl-EryACP6 with no detectable dehydration. These results establish the intrinsic syn dehydration stereochemistry and substrate diastereoselectivity of RifDH10 and highlight the critical role of the natural RifACP10 domain in chaperoning the proper recognition and processing of the natural ACP-bound undecaketide substrate. The 1.82 Å resolution structure of RifDH10 reveals the atomic-resolution details of the active site and allows modeling of the syn dehydration of the (2S,3S)-2-methyl-3-hydroxyacyl-RifACP10 substrate. These results suggest that generation of the characteristic cis double bond of the rifamycins occurs after formation of the full-length RifACP10-bound acyclic trans-unsaturated undecaketide intermediate, most likely during the subsequent macrolactamization catalyzed by the amide synthase RifF.
Organizational Affiliation:
Department of Chemistry and Biochemistry, The University of Texas at Austin , 1 University Station A5300, Austin, Texas 78712-0165, United States.