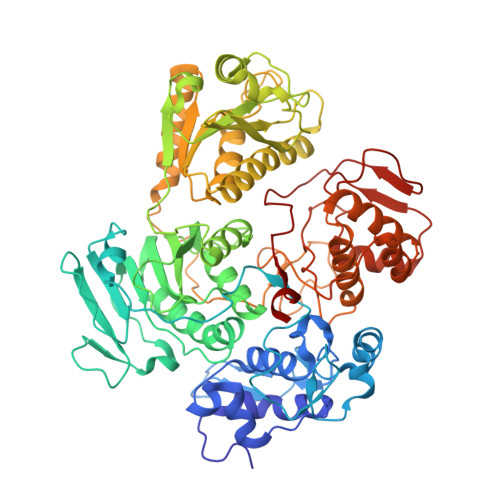
Structural and functional insights into alphavirus polyprotein processing and pathogenesis.
Shin, G., Yost, S.A., Miller, M.T., Elrod, E.J., Grakoui, A., Marcotrigiano, J.(2012) Proc Natl Acad Sci U S A 109: 16534-16539
- PubMed: 23010928
- DOI: https://doi.org/10.1073/pnas.1210418109
- Primary Citation of Related Structures:
4GUA - PubMed Abstract:
Alphaviruses, a group of positive-sense RNA viruses, are globally distributed arboviruses capable of causing rash, arthritis, encephalitis, and death in humans. The viral replication machinery consists of four nonstructural proteins (nsP1-4) produced as a single polyprotein. Processing of the polyprotein occurs in a highly regulated manner, with cleavage at the P2/3 junction influencing RNA template use during genome replication. Here, we report the structure of P23 in a precleavage form. The proteins form an extensive interface and nsP3 creates a ring structure that encircles nsP2. The P2/3 cleavage site is located at the base of a narrow cleft and is not readily accessible, suggesting a highly regulated cleavage. The nsP2 protease active site is over 40 Å away from the P2/3 cleavage site, supporting a trans cleavage mechanism. nsP3 contains a previously uncharacterized protein fold with a zinc-coordination site. Known mutations in nsP2 that result in formation of noncytopathic viruses or a temperature sensitive phenotype cluster at the nsP2/nsP3 interface. Structure-based mutations in nsP3 opposite the location of the nsP2 noncytopathic mutations prevent efficient cleavage of P23, affect RNA infectivity, and alter viral RNA production levels, highlighting the importance of the nsP2/nsP3 interaction in pathogenesis. A potential RNA-binding surface, spanning both nsP2 and nsP3, is proposed based on the location of ion-binding sites and adaptive mutations. These results offer unexpected insights into viral protein processing and pathogenesis that may be applicable to other polyprotein-encoding viruses such as HIV, hepatitis C virus (HCV), and Dengue virus.
Organizational Affiliation:
Center for Advanced Biotechnology and Medicine, Department of Chemistry and Chemical Biology, Rutgers University, Piscataway, NJ 08854, USA.