A bulky hydrophobic residue is not required to maintain the v-conformation of enzyme-bound thiamin diphosphate.
Andrews, F.H., Tom, A.R., Gunderman, P.R., Novak, W.R., McLeish, M.J.(2013) Biochemistry 52: 3028-3030
- PubMed: 23607689
- DOI: https://doi.org/10.1021/bi400368j
- Primary Citation of Related Structures:
4GG1, 4GM0, 4GM1, 4GM4, 4GP9, 4GPE, 4JD5 - PubMed Abstract:
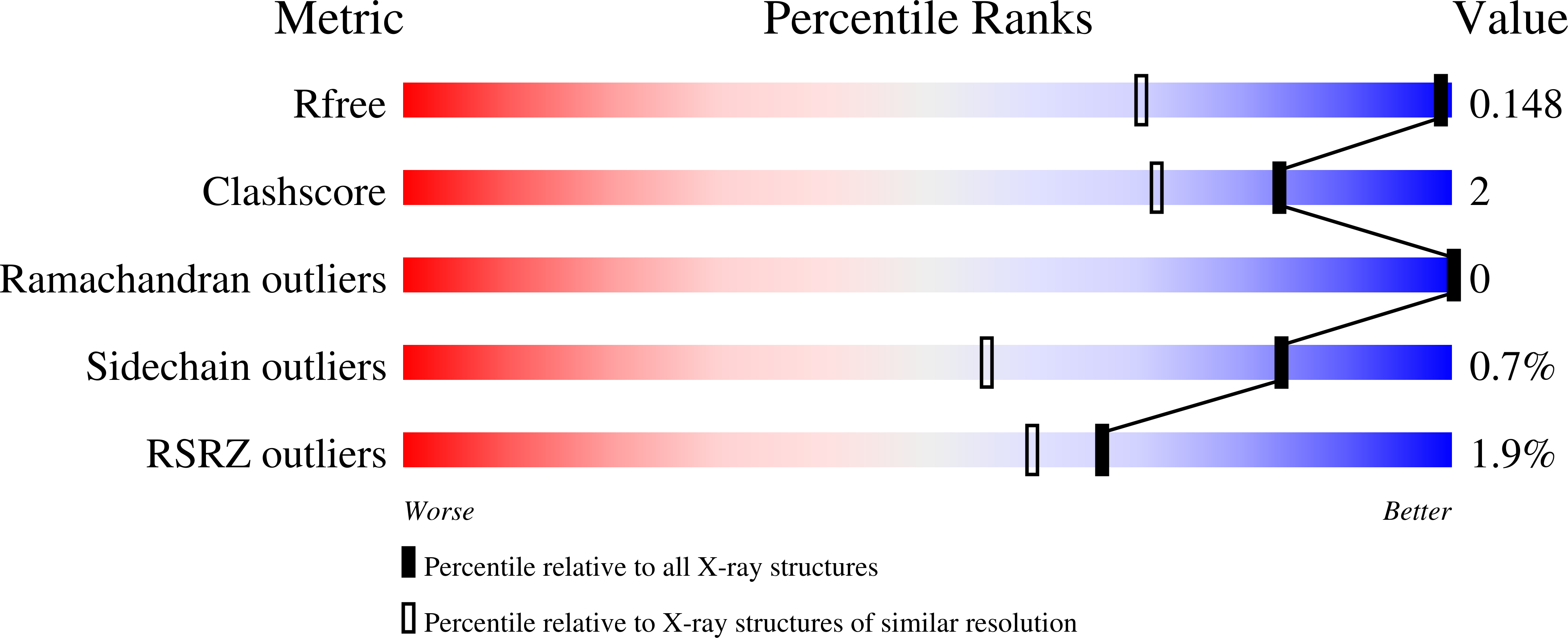
It is widely accepted that, in thiamin diphosphate (ThDP)-dependent enzymes, much of the rate acceleration is provided by the cofactor. Inter alia, the reactive conformation of ThDP, known as the V-conformation, has been attributed to the presence of a bulky hydrophobic residue located directly below the cofactor. Here we report the use of site-saturation mutagenesis to generate variants of this residue (Leu403) in benzoylformate decarboxylase. The observed 3 orders of magnitude range in k(cat)/K(m) values suggested that conformational changes in the cofactor could be influencing catalysis. However, X-ray structures of several variants were determined, and there was remarkably little change in ThDP conformation. Rather, it seemed that, once the V-conformation was attained, residue size and hydrophobicity were more important for enzyme activity.
Organizational Affiliation:
Department of Chemistry and Chemical Biology, Indiana University-Purdue University Indianapolis, Indianapolis, IN 46202, USA.