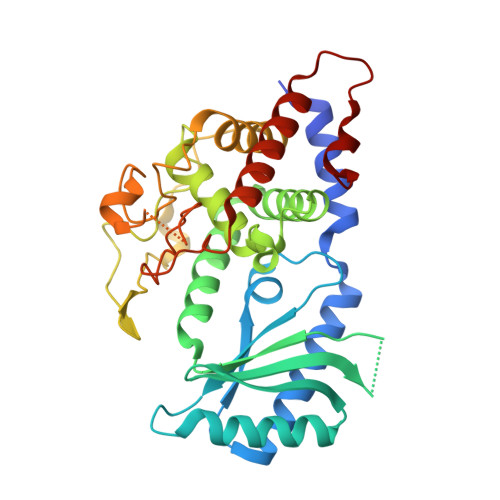
Crystal structures of the Cid1 poly (U) polymerase reveal the mechanism for UTP selectivity.
Lunde, B.M., Magler, I., Meinhart, A.(2012) Nucleic Acids Res 40: 9815-9824
- PubMed: 22885303
- DOI: https://doi.org/10.1093/nar/gks740
- Primary Citation of Related Structures:
4FH3, 4FH5, 4FHP, 4FHV, 4FHW, 4FHX, 4FHY - PubMed Abstract:
Polyuridylation is emerging as a ubiquitous post-translational modification with important roles in multiple aspects of RNA metabolism. These poly (U) tails are added by poly (U) polymerases with homology to poly (A) polymerases; nevertheless, the selection for UTP over ATP remains enigmatic. We report the structures of poly (U) polymerase Cid1 from Schizoscaccharomyces pombe alone and in complex with UTP, CTP, GTP and 3'-dATP. These structures reveal that each of the 4 nt can be accommodated at the active site; however, differences exist that suggest how the polymerase selects UTP over the other nucleotides. Furthermore, we find that Cid1 shares a number of common UTP recognition features with the kinetoplastid terminal uridyltransferases. Kinetic analysis of Cid1's activity for its preferred substrates, UTP and ATP, reveal a clear preference for UTP over ATP. Ultimately, we show that a single histidine in the active site plays a pivotal role for poly (U) activity. Notably, this residue is typically replaced by an asparagine residue in Cid1-family poly (A) polymerases. By mutating this histidine to an asparagine residue in Cid1, we diminished Cid1's activity for UTP addition and improved ATP incorporation, supporting that this residue is important for UTP selectivity.
Organizational Affiliation:
Department of Biomolecular Mechanisms, Max-Planck-Institute for Medical Research, Jahnstrasse 29, 69120 Heidelberg, Germany.