New 2-(Aryloxy)-3-phenylpropanoic Acids as Peroxisome Proliferator-Activated Receptor alpha/gamma Dual Agonists Able To Upregulate Mitochondrial Carnitine Shuttle System Gene Expression.
Laghezza, A., Pochetti, G., Lavecchia, A., Fracchiolla, G., Faliti, S., Piemontese, L., Di Giovanni, C., Iacobazzi, V., Infantino, V., Montanari, R., Capelli, D., Tortorella, P., Loiodice, F.(2013) J Med Chem 56: 60-72
- PubMed: 23171045
- DOI: https://doi.org/10.1021/jm301018z
- Primary Citation of Related Structures:
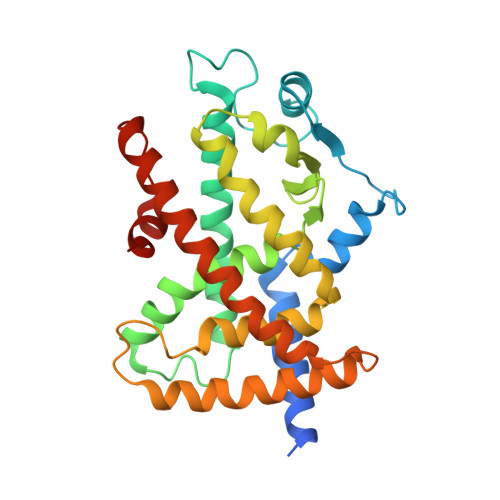
4E4K, 4E4Q - PubMed Abstract:
The preparation of a series of 2-(aryloxy)-3-phenylpropanoic acids, resulting from the introduction of different substituents into the biphenyl system of the previously reported peroxisome proliferator-activated receptor α/γ (PPARα/γ) dual agonist 1, allowed the identification of new ligands with higher potency on PPARα and fine-tuned moderate PPARγ activity. For the most promising stereoisomer (S)-16, X-ray and calorimetric studies in PPARγ revealed, at high ligand concentration, the presence of two molecules simultaneously bound to the receptor. On the basis of these results and docking experiments in both receptor subtypes, a molecular explanation was provided for its different behavior as a full and partial agonist of PPARα and PPARγ, respectively. The effects of (S)-16 on mitochondrial acylcarnitine carrier and carnitine-palmitoyl-transferase 1 gene expression, two key components of the carnitine shuttle system, were also investigated, allowing the hypothesis of a more beneficial pharmacological profile of this compound compared to the less potent PPARα agonist fibrates currently used in therapy.
Organizational Affiliation:
Dipartimento di Farmacia-Scienze del Farmaco and ‡Laboratorio di Biochimica e Biologia Molecolare, Dipartimento di Bioscienze, Biotecnologie e Biofarmaceutica, Università degli Studi di Bari "Aldo Moro", 70126 Bari, Italy.