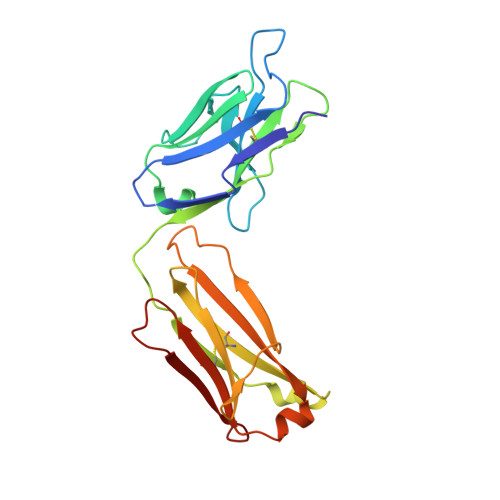
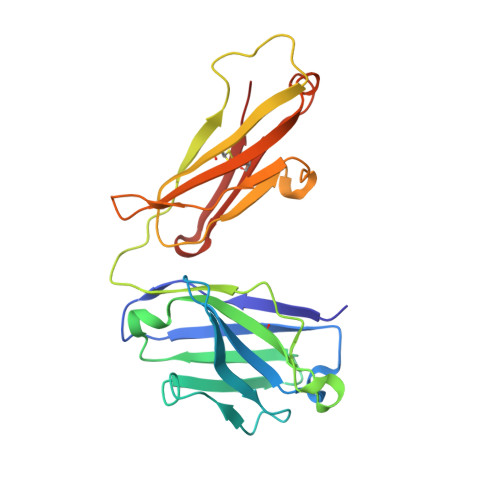
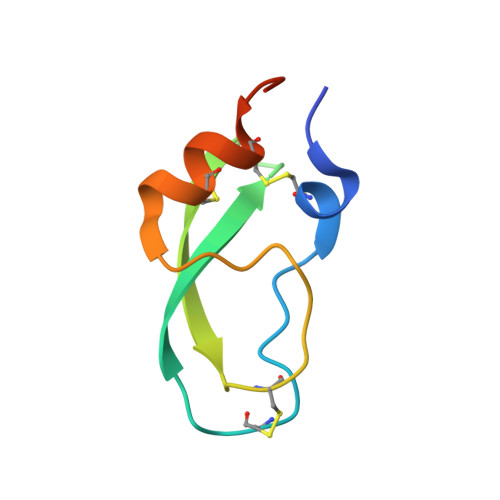
Hemostatic effect of a monoclonal antibody mAb 2021 blocking the interaction between FXa and TFPI in a rabbit hemophilia model.
Hilden, I., Lauritzen, B., Sorensen, B.B., Clausen, J.T., Jespersgaard, C., Krogh, B.O., Bowler, A.N., Breinholt, J., Gruhler, A., Svensson, L.A., Petersen, H.H., Petersen, L.C., Balling, K.W., Hansen, L., Hermit, M.B., Egebjerg, T., Friederichsen, B., Ezban, M., Bjorn, S.E.(2012) Blood 119: 5871-5878
- PubMed: 22563084
- DOI: https://doi.org/10.1182/blood-2012-01-401620
- Primary Citation of Related Structures:
4DTG - PubMed Abstract:
Hemophilia is treated by IV replacement therapy with Factor VIII (FVIII) or Factor IX (FIX), either on demand to resolve bleeding, or as prophylaxis. Improved treatment may be provided by drugs designed for subcutaneous and less frequent administration with a reduced risk of inhibitor formation. Tissue factor pathway inhibitor (TFPI) down-regulates the initiation of coagulation by inhibition of Factor VIIa (FVIIa)/tissue factor/Factor Xa (FVIIa/TF/FXa). Blockage of TFPI inhibition may facilitate thrombin generation in a hemophilic setting. A high-affinity (K(D) = 25pM) mAb, mAb 2021, against TFPI was investigated. Binding of mAb 2021 to TFPI effectively prevented inhibition of FVIIa/TF/FXa and improved clot formation in hemophilia blood and plasma. The binding epitope on the Kunitz-type protease inhibitor domain 2 of TFPI was mapped by crystallography, and showed an extensive overlap with the FXa contact region highlighting a structural basis for its mechanism of action. In a rabbit hemophilia model, an intravenous or subcutaneous dose significantly reduced cuticle bleeding. mAb 2021 showed an effect comparable with that of rFVIIa. Cuticle bleeding in the model was reduced for at least 7 days by a single intravenous dose of mAb 2021. This study suggests that neutralization of TFPI by mAb 2021 may constitute a novel treatment option in hemophilia.
Organizational Affiliation:
Biopharmaceuticals Research Unit, Novo Nordisk, Maaloev, Denmark. IDAH@novonordisk.com