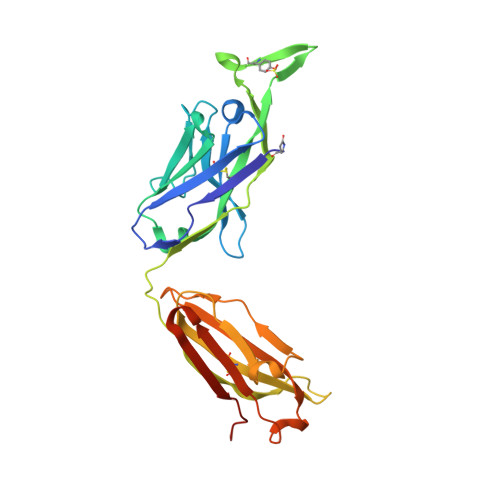
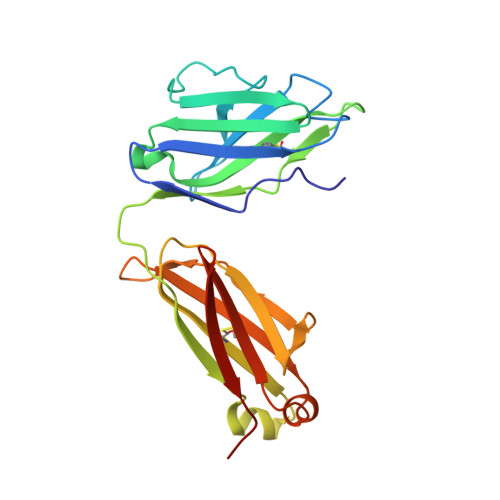
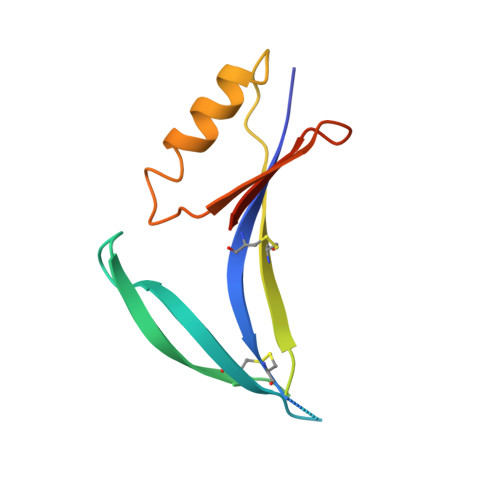
Structural basis for diverse N-glycan recognition by HIV-1-neutralizing V1-V2-directed antibody PG16.
Pancera, M., Shahzad-Ul-Hussan, S., Doria-Rose, N.A., McLellan, J.S., Bailer, R.T., Dai, K., Loesgen, S., Louder, M.K., Staupe, R.P., Yang, Y., Zhang, B., Parks, R., Eudailey, J., Lloyd, K.E., Blinn, J., Alam, S.M., Haynes, B.F., Amin, M.N., Wang, L.X., Burton, D.R., Koff, W.C., Nabel, G.J., Mascola, J.R., Bewley, C.A., Kwong, P.D.(2013) Nat Struct Mol Biol 20: 804-813
- PubMed: 23708607
- DOI: https://doi.org/10.1038/nsmb.2600
- Primary Citation of Related Structures:
4DQO - PubMed Abstract:
HIV-1 uses a diverse N-linked-glycan shield to evade recognition by antibody. Select human antibodies, such as the clonally related PG9 and PG16, recognize glycopeptide epitopes in the HIV-1 V1-V2 region and penetrate this shield, but their ability to accommodate diverse glycans is unclear. Here we report the structure of antibody PG16 bound to a scaffolded V1-V2, showing an epitope comprising both high mannose-type and complex-type N-linked glycans. We combined structure, NMR and mutagenesis analyses to characterize glycan recognition by PG9 and PG16. Three PG16-specific residues, arginine, serine and histidine (RSH), were critical for binding sialic acid on complex-type glycans, and introduction of these residues into PG9 produced a chimeric antibody with enhanced HIV-1 neutralization. Although HIV-1-glycan diversity facilitates evasion, antibody somatic diversity can overcome this and can provide clues to guide the design of modified antibodies with enhanced neutralization.
Organizational Affiliation:
Vaccine Research Center, National Institute of Allergy and Infectious Diseases, US National Institutes of Health, Bethesda, Maryland, USA.