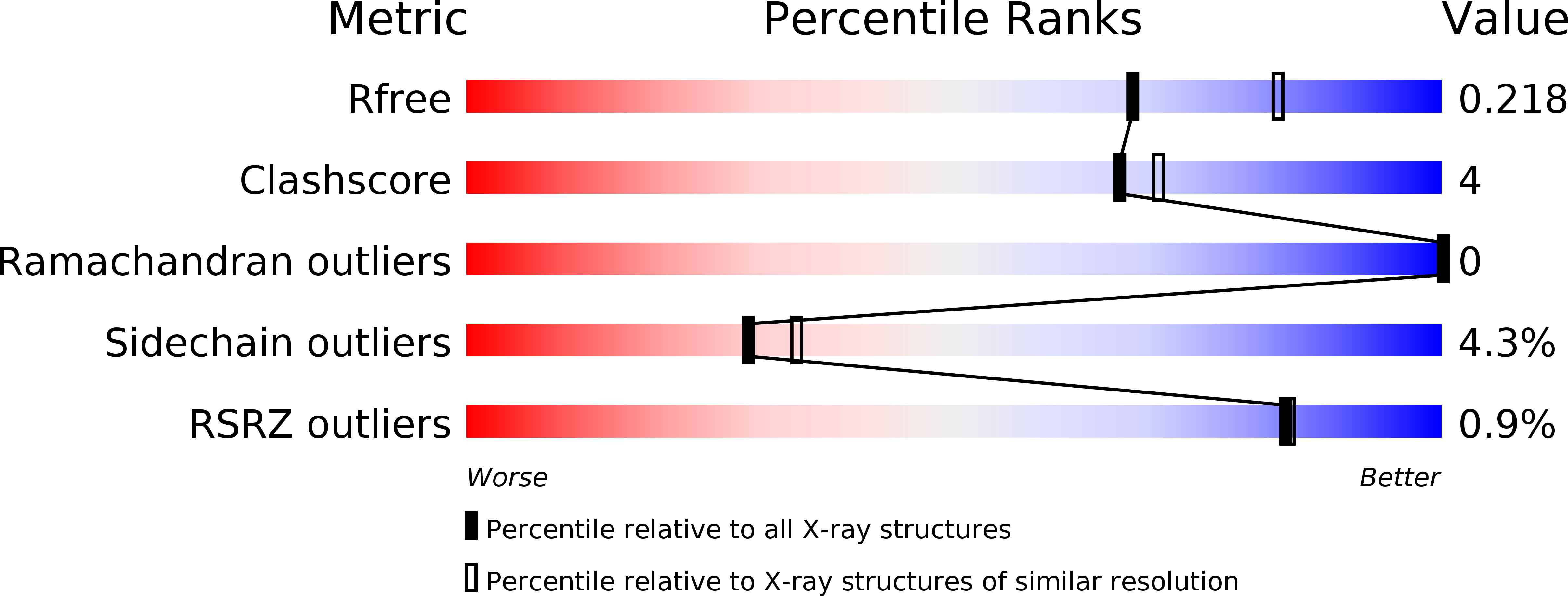
Crystal Structures and Functional Characterization of Wild-Type Cyp101D1 and its Active Site Mutants.
Batabyal, D., Poulos, T.L.(2013) Biochemistry 52: 8898
- PubMed: 24261604
- DOI: https://doi.org/10.1021/bi401330c
- Primary Citation of Related Structures:
4C9K, 4C9L, 4C9M, 4C9N, 4C9O, 4C9P - PubMed Abstract:
Although CYP101D1 and P450cam catalyze the same reaction at similar rates and share strikingly similar active site architectures, there are significant functional differences. CYP101D1 thus provides an opportunity to probe what structural and functional features must be shared and what features can differ but maintain the high catalytic efficiency. Crystal structures of the cyanide complex of wild-type CYP101D1 and it active site mutants, D259N and T260A, have been determined. The conformational changes in CYP101D1 upon cyanide binding are very similar to those of P450cam, indicating a similar mechanism for proton delivery during oxygen activation using solvent-assisted proton transfer. The D259N-CN- complex shows a perturbed solvent structure compared to that of the wild type, which is similar to what was observed in the oxy complex of the corresonding D251N mutant in P450cam. As in P450cam, the T260A mutant is highly uncoupled while the D259N mutant gives barely detectable activity. Despite these similarities, CYP101D1 is able to use the P450cam redox partners while P450cam cannot use the CYP101D1 redox partners. Thus, the strict requirement of P450cam for its own redox partner is relaxed in CYP101D1. Differences in the local environment of the essential Asp (Asp259 in CYP101D1) provide a strucutral basis for understanding these functional differences.
Organizational Affiliation:
Departments of Molecular Biology and Biochemistry, Chemistry, and Pharmaceutical Sciences, University of California , Irvine, California 92697-3900, United States.