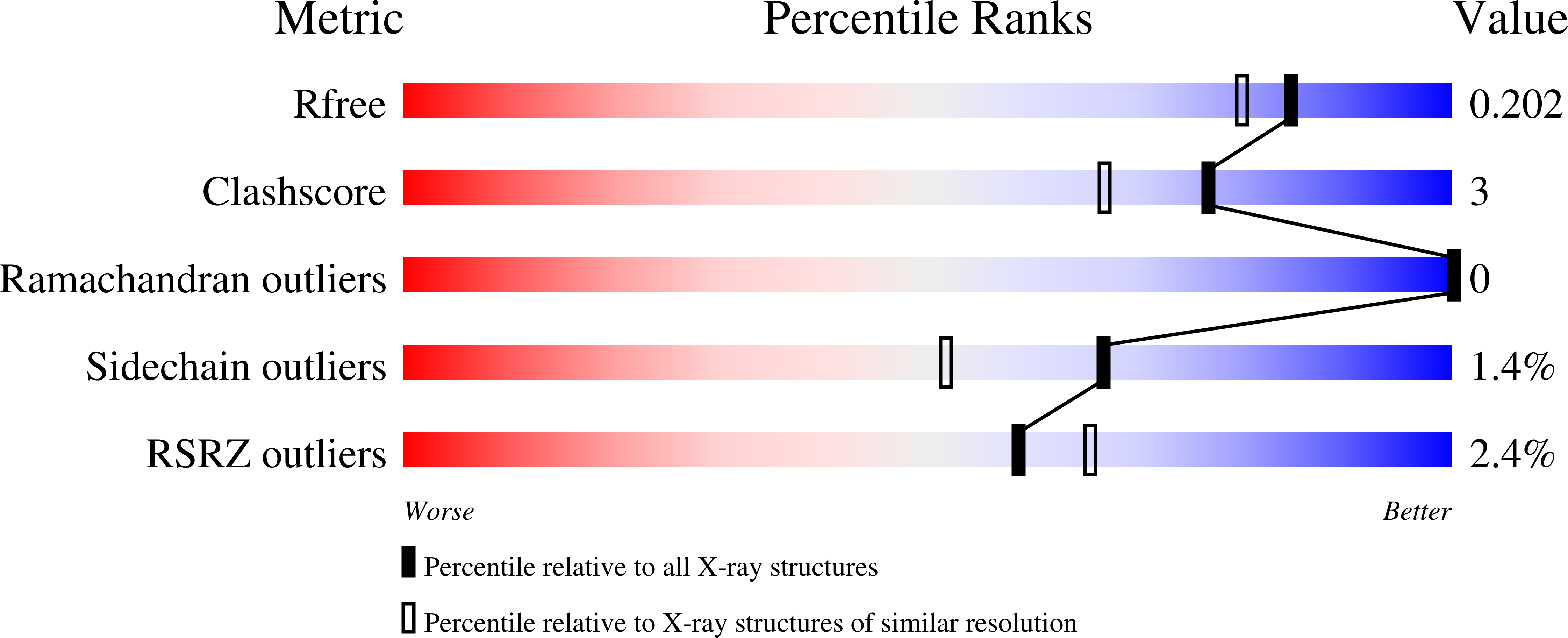
Correlating Efficacy and Desensitization with Gluk2 Ligand-Binding Domain Movements.
Nayeem, N., Mayans, O., Green, T.(2013) Open Biol 3: 0051
- PubMed: 23720540
- DOI: https://doi.org/10.1098/rsob.130051
- Primary Citation of Related Structures:
4BDL, 4BDM, 4BDN, 4BDO, 4BDQ, 4BDR - PubMed Abstract:
Gating of AMPA- and kainate-selective ionotropic glutamate receptors can be defined in terms of ligand affinity, efficacy and the rate and extent of desensitization. Crucial insights into all three elements have come from structural studies of the ligand-binding domain (LBD). In particular, binding-cleft closure is associated with efficacy, whereas dissociation of the dimer formed by neighbouring LBDs is linked with desensitization. We have explored these relationships in the kainate-selective subunit GluK2 by studying the effects of mutating two residues (K531 and R775) that form key contacts within the LBD dimer interface, but whose truncation unexpectedly attenuates desensitization. One mutation (K531A) also switches the relative efficacies of glutamate and kainate. LBD crystal structures incorporating these mutations revealed several conformational changes that together explain their phenotypes. K531 truncation results in new dimer contacts, consistent with slower desensitization and sideways movement in the ligand-binding cleft correlating with efficacy. The tested mutants also disrupted anion binding; no chloride was detected in the dimer-interface site, including in R775A where absence of chloride was the only structural change evident. From this, we propose that the charge balance in the GluK2 LBD dimer interface maintains a degree of instability, necessary for rapid and complete desensitization.
Organizational Affiliation:
Department of Pharmacology, University of Liverpool, Liverpool L69 3GE, UK.