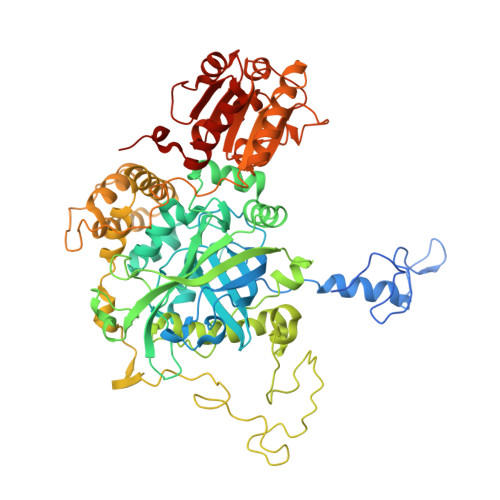
Structure, Recombinant Expression and Mutagenesis Studies of the Catalase with Oxidase Activity from Scytalidium Thermophilum
Yuzugullu, Y., Trinh, C.H., Smith, M.A., Pearson, A.R., Phillips, S.E.V., Sutay Kocabas, D., Bakir, U., Ogel, Z.B., McPherson, M.J.(2013) Acta Crystallogr D Biol Crystallogr 69: 398
- PubMed: 23519415
- DOI: https://doi.org/10.1107/S0907444912049001
- Primary Citation of Related Structures:
4AUE, 4AUL, 4AUM, 4AUN - PubMed Abstract:
Scytalidium thermophilum produces a catalase with phenol oxidase activity (CATPO) that catalyses the decomposition of hydrogen peroxide into oxygen and water and also oxidizes various phenolic compounds. A codon-optimized catpo gene was cloned and expressed in Escherichia coli. The crystal structures of native and recombinant S. thermophilum CATPO and two variants, H82N and V123F, were determined at resolutions of 2.7, 1.4, 1.5 and 1.9 Å, respectively. The structure of CATPO reveals a homotetramer with 698 residues per subunit and with strong structural similarity to Penicillium vitale catalase. The haem component is cis-hydroxychlorin γ-spirolactone, which is rotated 180° with respect to small-subunit catalases. The haem-binding pocket contains two highly conserved water molecules on the distal side. The H82N mutation resulted in conversion of the native d-type haem to a b-type haem. Kinetic studies of the H82N and V123F mutants indicate that both activities are likely to be associated with the haem centre and suggest that the secondary oxidase activity may be a general feature of catalases in the absence of hydrogen peroxide.
Organizational Affiliation:
Department of Biology, Kocaeli University, 41380 Kocaeli, Turkey.