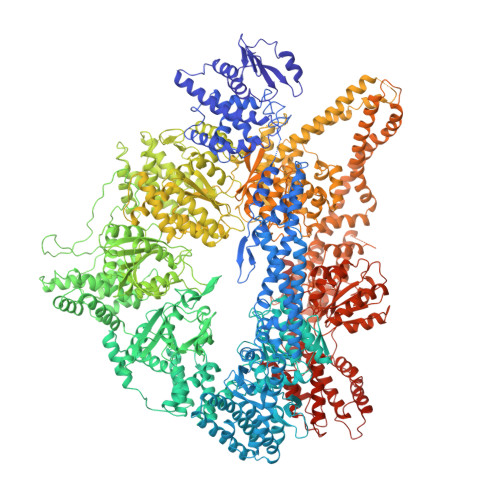
Insights Into Dynein Motor Domain Function from a 3.3 Angstrom Crystal Structure
Schmidt, H., Gleave, E.S., Carter, A.P.(2012) Nat Struct Mol Biol 19: 492
- PubMed: 22426545
- DOI: https://doi.org/10.1038/nsmb.2272
- Primary Citation of Related Structures:
4AI6, 4AKG, 4AKH, 4AKI - PubMed Abstract:
Dyneins power the beating of cilia and flagella, transport various intracellular cargos and are necessary for mitosis. All dyneins have a ∼300-kDa motor domain consisting of a ring of six AAA+ domains. ATP hydrolysis in the AAA+ ring drives the cyclic relocation of a motile element, the linker domain, to generate the force necessary for movement. How the linker interacts with the ring during the ATP hydrolysis cycle is not known. Here we present a 3.3-Å crystal structure of the motor domain of Saccharomyces cerevisiae cytoplasmic dynein, crystallized in the absence of nucleotides. The linker is docked to a conserved site on AAA5, which is confirmed by mutagenesis as functionally necessary. Nucleotide soaking experiments show that the main ATP hydrolysis site in dynein (AAA1) is in a low-nucleotide affinity conformation and reveal the nucleotide interactions of the other three sites (AAA2, AAA3 and AAA4).
Organizational Affiliation:
Medical Research Council Laboratory of Molecular Biology, Cambridge, UK.