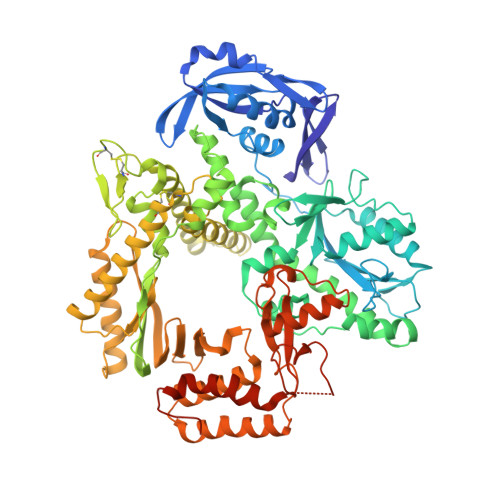
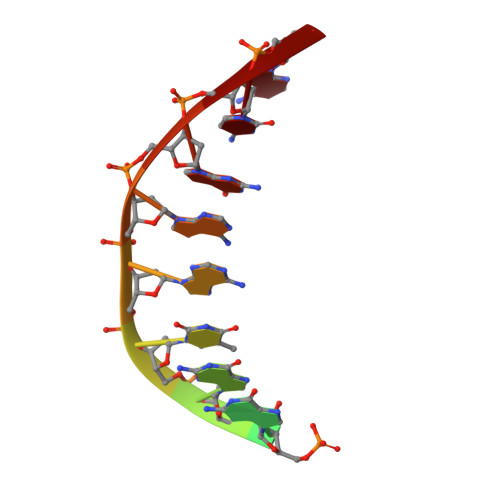
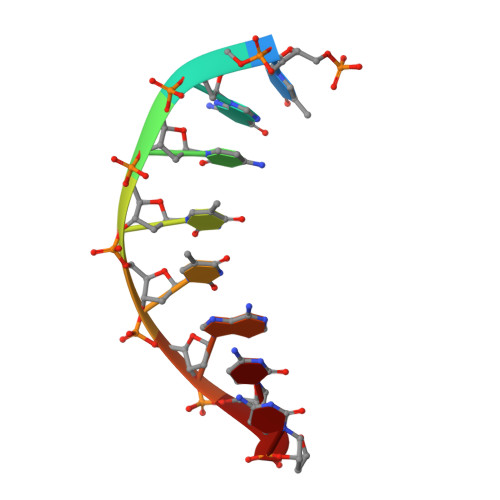
Structures of an Apo and a Binary Complex of an Evolved Archeal B Family DNA Polymerase Capable of Synthesising Highly Cy-Dye Labelled DNA.
Wynne, S.A., Pinheiro, V.B., Holliger, P., Leslie, A.G.(2013) PLoS One 8: 70892
- PubMed: 23940661
- DOI: https://doi.org/10.1371/journal.pone.0070892
- Primary Citation of Related Structures:
4AHC, 4AIL - PubMed Abstract:
Thermophilic DNA polymerases of the polB family are of great importance in biotechnological applications including high-fidelity PCR. Of particular interest is the relative promiscuity of engineered versions of the exo- form of polymerases from the Thermo- and Pyrococcales families towards non-canonical substrates, which enables key advances in Next-generation sequencing. Despite this there is a paucity of structural information to guide further engineering of this group of polymerases. Here we report two structures, of the apo form and of a binary complex of a previously described variant (E10) of Pyrococcus furiosus (Pfu) polymerase with an ability to fully replace dCTP with Cyanine dye-labeled dCTP (Cy3-dCTP or Cy5-dCTP) in PCR and synthesise highly fluorescent "CyDNA" densely decorated with cyanine dye heterocycles. The apo form of Pfu-E10 closely matches reported apo form structures of wild-type Pfu. In contrast, the binary complex (in the replicative state with a duplex DNA oligonucleotide) reveals a closing movement of the thumb domain, increasing the contact surface with the nascent DNA duplex strand. Modelling based on the binary complex suggests how bulky fluorophores may be accommodated during processive synthesis and has aided the identification of residues important for the synthesis of unnatural nucleic acid polymers.
Organizational Affiliation:
Medical Research Council Laboratory of Molecular Biology, Cambridge, United Kingdom.