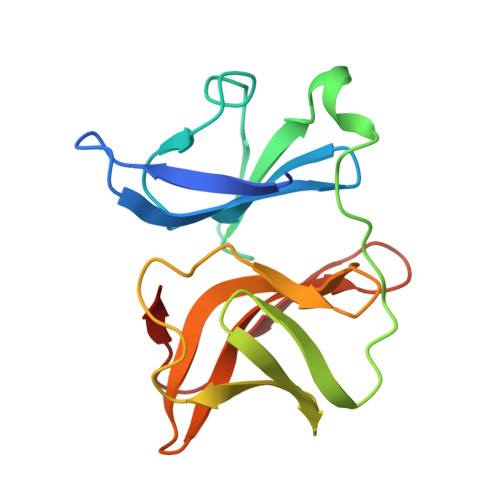
Crystal Structure of Aura Virus Capsid Protease and its Complex with Dioxane: New Insights Into Capsid-Glycoprotein Molecular Contacts.
Aggarwal, M., Tapas, S., Preeti, Siwach, A., Kumar, P., Kuhn, R.J., Tomar, S.(2012) PLoS One 7: 51288
- PubMed: 23251484
- DOI: https://doi.org/10.1371/journal.pone.0051288
- Primary Citation of Related Structures:
4AGJ, 4AGK - PubMed Abstract:
The nucleocapsid core interaction with endodomains of glycoproteins plays a critical role in the alphavirus life cycle that is essential to virus budding. Recent cryo-electron microscopy (cryo-EM) studies provide structural insights into key interactions between capsid protein (CP) and trans-membrane glycoproteins E1 and E2. CP possesses a chymotrypsin-like fold with a hydrophobic pocket at the surface responsible for interaction with glycoproteins. In the present study, crystal structures of the protease domain of CP from Aura virus and its complex with dioxane were determined at 1.81 and 1.98 Å resolution respectively. Due to the absence of crystal structures, homology models of E1 and E2 from Aura virus were generated. The crystal structure of CP and structural models of E1 and E2 were fitted into the cryo-EM density map of Venezuelan equine encephalitis virus (VEEV) for detailed analysis of CP-glycoprotein interactions. Structural analysis revealed that the E2 endodomain consists of a helix-loop-helix motif where the loop region fits into the hydrophobic pocket of CP. Our studies suggest that Cys397, Cys418 and Tyr401 residues of E2 are involved in stabilizing the structure of E2 endodomain. Density map fitting analysis revealed that Pro405, a conserved E2 residue is present in the loop region of the E2 endodomain helix-loop-helix structure and makes intermolecular hydrophobic contacts with the capsid. In the Aura virus capsid protease (AVCP)-dioxane complex structure, dioxane occupies the hydrophobic pocket on CP and structurally mimics the hydrophobic pyrollidine ring of Pro405 in the loop region of E2.
Organizational Affiliation:
Department of Biotechnology, Indian Institute of Technology, Roorkee, Roorkee, India.