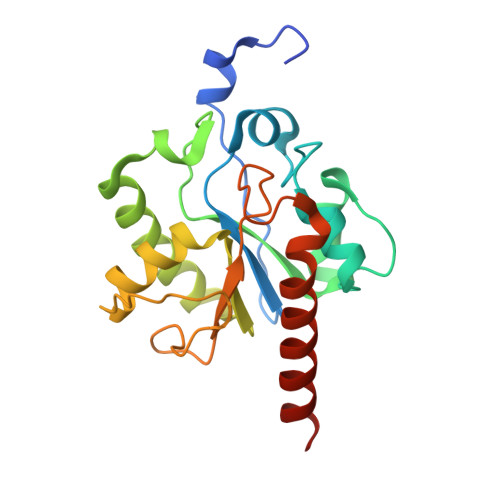
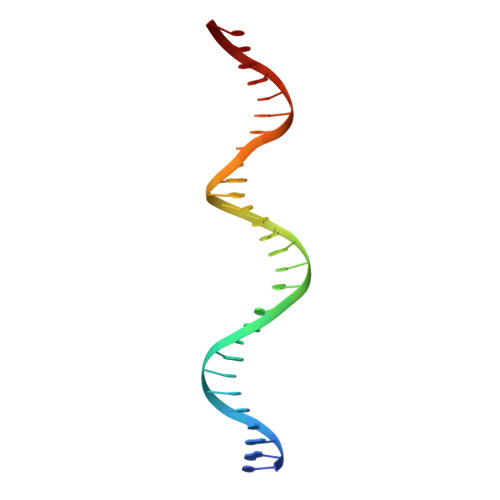
Thymine DNA glycosylase exhibits negligible affinity for nucleobases that it removes from DNA.
Malik, S.S., Coey, C.T., Varney, K.M., Pozharski, E., Drohat, A.C.(2015) Nucleic Acids Res 43: 9541-9552
- PubMed: 26358812
- DOI: https://doi.org/10.1093/nar/gkv890
- Primary Citation of Related Structures:
4XEG, 4Z3A, 4Z47, 4Z7B, 4Z7Z - PubMed Abstract:
Thymine DNA Glycosylase (TDG) performs essential functions in maintaining genetic integrity and epigenetic regulation. Initiating base excision repair, TDG removes thymine from mutagenic G ·: T mispairs caused by 5-methylcytosine (mC) deamination and other lesions including uracil (U) and 5-hydroxymethyluracil (hmU). In DNA demethylation, TDG excises 5-formylcytosine (fC) and 5-carboxylcytosine (caC), which are generated from mC by Tet (ten-eleven translocation) enzymes. Using improved crystallization conditions, we solved high-resolution (up to 1.45 Å) structures of TDG enzyme-product complexes generated from substrates including G·U, G·T, G·hmU, G·fC and G·caC. The structures reveal many new features, including key water-mediated enzyme-substrate interactions. Together with nuclear magnetic resonance experiments, the structures demonstrate that TDG releases the excised base from its tight product complex with abasic DNA, contrary to previous reports. Moreover, DNA-free TDG exhibits no significant binding to free nucleobases (U, T, hmU), indicating a Kd >> 10 mM. The structures reveal a solvent-filled channel to the active site, which might facilitate dissociation of the excised base and enable caC excision, which involves solvent-mediated acid catalysis. Dissociation of the excised base allows TDG to bind the beta rather than the alpha anomer of the abasic sugar, which might stabilize the enzyme-product complex.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of Maryland School of Medicine, Baltimore, MD 21201, USA.