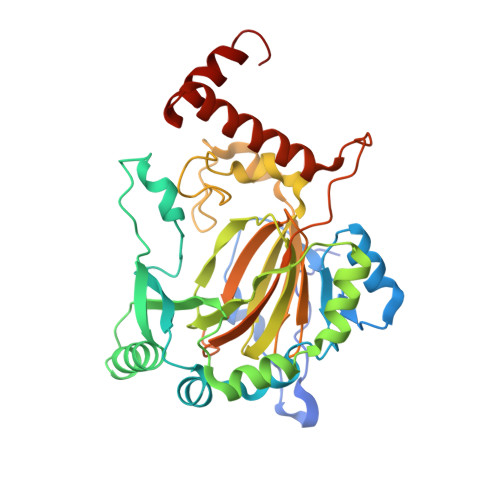
Substrate Promotes Productive Gas Binding in the alpha-Ketoglutarate-Dependent Oxygenase FIH.
Taabazuing, C.Y., Fermann, J., Garman, S., Knapp, M.J.(2016) Biochemistry 55: 277-286
- PubMed: 26727884
- DOI: https://doi.org/10.1021/acs.biochem.5b01003
- Primary Citation of Related Structures:
4Z1V, 4Z2W - PubMed Abstract:
The Fe(2+)/α-ketoglutarate (αKG)-dependent oxygenases use molecular oxygen to conduct a wide variety of reactions with important biological implications, such as DNA base excision repair, histone demethylation, and the cellular hypoxia response. These enzymes follow a sequential mechanism in which O2 binds and reacts after the primary substrate binds, making those structural factors that promote productive O2 binding central to their chemistry. A large challenge in this field is to identify strategies that engender productive turnover. Factor inhibiting HIF (FIH) is a Fe(2+)/αKG-dependent oxygenase that forms part of the O2 sensing machinery in human cells by hydroxylating the C-terminal transactivation domain (CTAD) found within the HIF-1α protein. The structure of FIH was determined with the O2 analogue NO bound to Fe, offering the first direct insight into the gas binding geometry in this enzyme. Through a combination of density functional theory calculations, {FeNO}(7) electron paramagnetic resonance spectroscopy, and ultraviolet-visible absorption spectroscopy, we demonstrate that CTAD binding stimulates O2 reactivity by altering the orientation of the bound gas molecule. Although unliganded FIH binds NO with moderate affinity, the bound gas can adopt either of two orientations with similar stability; upon CTAD binding, NO adopts a single preferred orientation that is appropriate for supporting oxidative decarboxylation. Combined with other studies of related enzymes, our data suggest that substrate-induced reorientation of bound O2 is the mechanism utilized by the αKG oxygenases to tightly couple O2 activation to substrate hydroxylation.
Organizational Affiliation:
Department of Chemistry, University of Massachusetts , Amherst, Massachusetts 01003, United States.