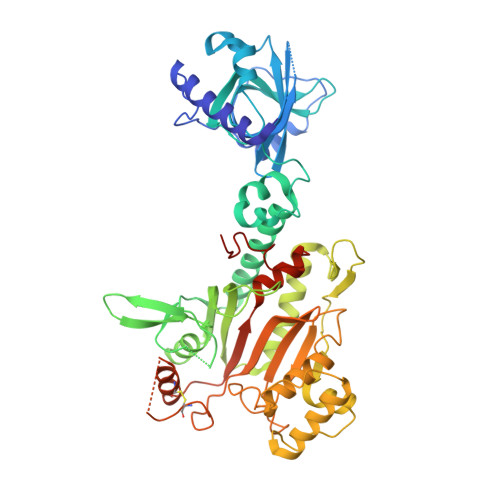
Structural Basis for Specific Inhibition of tRNA Synthetase by an ATP Competitive Inhibitor.
Fang, P., Han, H., Wang, J., Chen, K., Chen, X., Guo, M.(2015) Chem Biol 22: 734-744
- PubMed: 26074468
- DOI: https://doi.org/10.1016/j.chembiol.2015.05.007
- Primary Citation of Related Structures:
4YCU, 4YCV - PubMed Abstract:
Pharmaceutical inhibitors of aminoacyl-tRNA synthetases demand high species and family specificity. The antimalarial ATP-mimetic cladosporin selectively inhibits Plasmodium falciparum LysRS (PfLysRS). How the binding to a universal ATP site achieves the specificity is unknown. Here we report three crystal structures of cladosporin with human LysRS, PfLysRS, and a Pf-like human LysRS mutant. In all three structures, cladosporin occupies the class defining ATP-binding pocket, replacing the adenosine portion of ATP. Three residues holding the methyltetrahydropyran moiety of cladosporin are critical for the specificity of cladosporin against LysRS over other class II tRNA synthetase families. The species-exclusive inhibition of PfLysRS is linked to a structural divergence beyond the active site that mounts a lysine-specific stabilizing response to binding cladosporin. These analyses reveal that inherent divergence of tRNA synthetase structural assembly may allow for highly specific inhibition even through the otherwise universal substrate binding pocket and highlight the potential for structure-driven drug development.
Organizational Affiliation:
Department of Cancer Biology, The Scripps Research Institute, Scripps Florida, 130 Scripps Way, Jupiter, FL 33458, USA. Electronic address: pfang@scripps.edu.