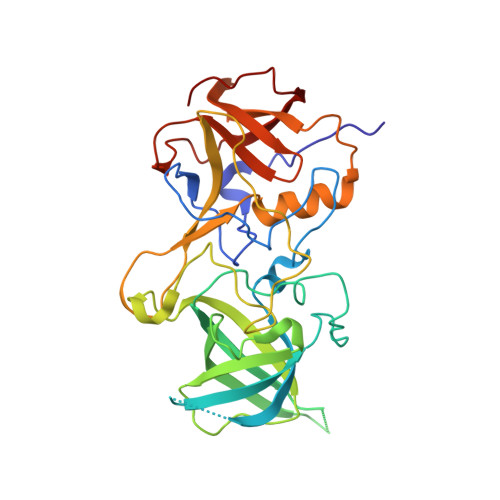
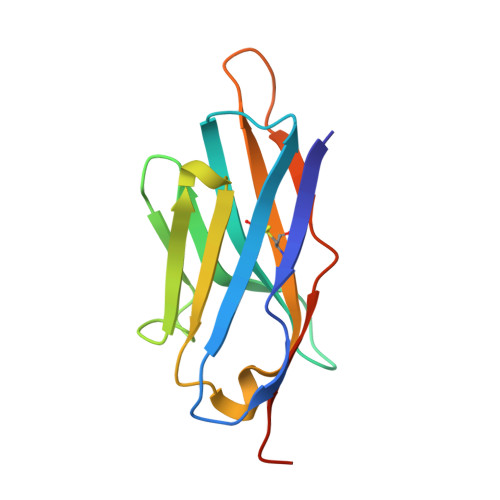
Nanobody binding to a conserved epitope promotes norovirus particle disassembly.
Koromyslova, A.D., Hansman, G.S.(2015) J Virol 89: 2718-2730
- PubMed: 25520510
- DOI: https://doi.org/10.1128/JVI.03176-14
- Primary Citation of Related Structures:
4X7C, 4X7D, 4X7E, 4X7F - PubMed Abstract:
Human noroviruses are icosahedral single-stranded RNA viruses. The capsid protein is divided into shell (S) and protruding (P) domains, which are connected by a flexible hinge region. There are numerous genetically and antigenically distinct noroviruses, and the dominant strains evolve every other year. Vaccine and antiviral development is hampered by the difficulties in growing human norovirus in cell culture and the continually evolving strains. Here, we show the X-ray crystal structures of human norovirus P domains in complex with two different nanobodies. One nanobody, Nano-85, was broadly reactive, while the other, Nano-25, was strain specific. We showed that both nanobodies bound to the lower region on the P domain and had nanomolar affinities. The Nano-85 binding site mainly comprised highly conserved amino acids among the genetically distinct genogroup II noroviruses. Several of the conserved residues also were recognized by a broadly reactive monoclonal antibody, which suggested this region contained a dominant epitope. Superposition of the P domain nanobody complex structures into a cryoelectron microscopy particle structure revealed that both nanobodies bound at occluded sites on the particles. The flexible hinge region, which contained ~10 to 12 amino acids, likely permitted a certain degree of P domain movement on the particles in order to accommodate the nanobodies. Interestingly, the Nano-85 binding interaction with intact particles caused the particles to disassemble in vitro. Altogether, these results suggested that the highly conserved Nano-85 binding epitope contained a trigger mechanism for particle disassembly. Principally, this epitope represents a potential site of norovirus vulnerability. We characterized two different nanobodies (Nano-85 and Nano-25) that bind to human noroviruses. Both nanobodies bound with high affinities to the lower region of the P domain, which was occluded on intact particles. Nano-25 was specific for GII.10, whereas Nano-85 bound several different GII genotypes, including GII.4, GII.10, and GII.12. We showed that Nano-85 was able to detect norovirus virions in clinical stool specimens using a sandwich enzyme-linked immunosorbent assay. Importantly, we found that Nano-85 binding to intact particles caused the particles to disassemble. We believe that with further testing, Nano-85 not only will work as a diagnostic reagent in norovirus detection systems but also could function as a broadly reactive GII norovirus antiviral.
Organizational Affiliation:
Schaller Research Group at the University of Heidelberg and the DKFZ, Germany, Heidelberg, Germany, and Department of Infectious Diseases, Virology, University of Heidelberg, Germany, Heidelberg, Germany.