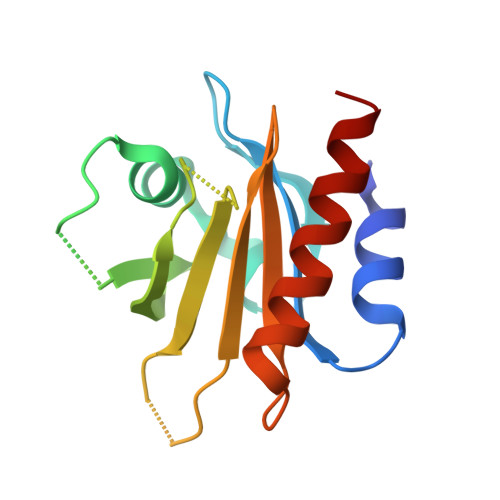
Structural basis for mutation-induced destabilization of profilin 1 in ALS.
Boopathy, S., Silvas, T.V., Tischbein, M., Jansen, S., Shandilya, S.M., Zitzewitz, J.A., Landers, J.E., Goode, B.L., Schiffer, C.A., Bosco, D.A.(2015) Proc Natl Acad Sci U S A 112: 7984-7989
- PubMed: 26056300
- DOI: https://doi.org/10.1073/pnas.1424108112
- Primary Citation of Related Structures:
4X1L, 4X1M, 4X25 - PubMed Abstract:
Mutations in profilin 1 (PFN1) are associated with amyotrophic lateral sclerosis (ALS); however, the pathological mechanism of PFN1 in this fatal disease is unknown. We demonstrate that ALS-linked mutations severely destabilize the native conformation of PFN1 in vitro and cause accelerated turnover of the PFN1 protein in cells. This mutation-induced destabilization can account for the high propensity of ALS-linked variants to aggregate and also provides rationale for their reported loss-of-function phenotypes in cell-based assays. The source of this destabilization is illuminated by the X-ray crystal structures of several PFN1 proteins, revealing an expanded cavity near the protein core of the destabilized M114T variant. In contrast, the E117G mutation only modestly perturbs the structure and stability of PFN1, an observation that reconciles the occurrence of this mutation in the control population. These findings suggest that a destabilized form of PFN1 underlies PFN1-mediated ALS pathogenesis.
Organizational Affiliation:
Department of Neurology, University of Massachusetts Medical School, Worcester, MA 01605;