Design and Synthesis of Highly Potent and Isoform Selective JNK3 Inhibitors: SAR Studies on Aminopyrazole Derivatives.
Zheng, K., Iqbal, S., Hernandez, P., Park, H., LoGrasso, P.V., Feng, Y.(2014) J Med Chem 57: 10013-10030
- PubMed: 25393557
- DOI: https://doi.org/10.1021/jm501256y
- Primary Citation of Related Structures:
4WHZ - PubMed Abstract:
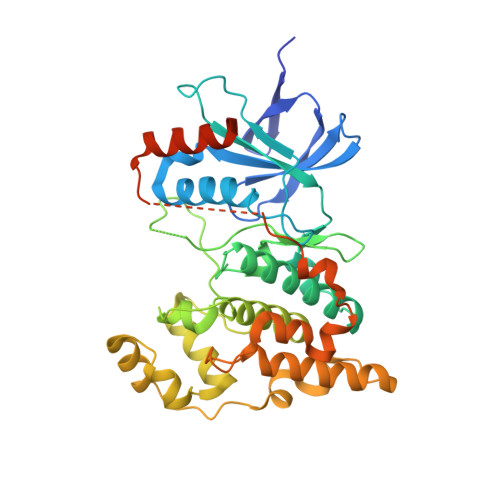
The c-jun N-terminal kinase 3 (JNK3) is expressed primarily in the brain. Numerous reports have shown that inhibition of JNK3 is a promising strategy for treatment of neurodegeneration. The optimization of aminopyrazole-based JNK3 inhibitors with improved potency, isoform selectivity, and pharmacological properties by structure-activity relationship (SAR) studies utilizing biochemical and cell-based assays, and structure-based drug design is reported. These inhibitors had high selectivity over JNK1 and p38α, minimal cytotoxicity, potent inhibition of 6-OHDA-induced mitochondrial membrane potential dissipation and ROS generation, and good drug metabolism and pharmacokinetic (DMPK) properties for iv dosing. 26n was profiled against 464 kinases and was found to be highly selective hitting only seven kinases with >80% inhibition at 10 μM. Moreover, 26n showed good solubility, good brain penetration, and good DMPK properties. Finally, the crystal structure of 26k in complex with JNK3 was solved at 1.8 Å to explore the binding mode of aminopyrazole based JNK3 inhibitors.
Organizational Affiliation:
Medicinal Chemistry, ‡Discovery Biology, §Crystallography/Modeling Facility, Translational Research Institute, and ∥Department of Molecular Therapeutics, Scripps Florida, The Scripps Research Institute , 130 Scripps Way, No. 2A1, Jupiter, Florida 33458, United States.