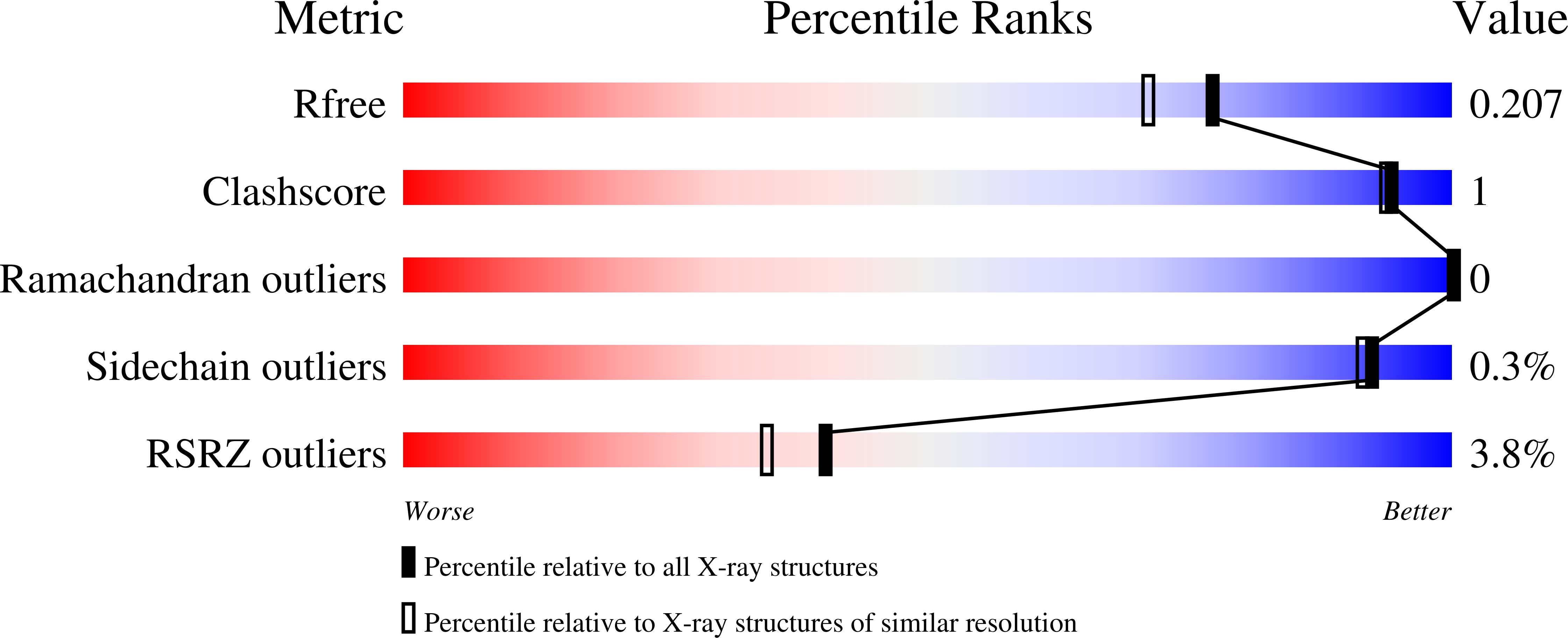
Structure-Based Design, Synthesis and Evaluation in Vitro of Arylnaphthyridinones, Arylpyridopyrimidinones and Their Tetrahydro Derivatives as Inhibitors of the Tankyrases.
Kumpan, K., Nathubhai, A., Zhang, C., Wood, P.J., Lloyd, M.D., Thompson, A.S., Haikarainen, T., Lehtio, L., Threadgill, M.D.(2015) Bioorg Med Chem 23: 3013
- PubMed: 26026769
- DOI: https://doi.org/10.1016/j.bmc.2015.05.005
- Primary Citation of Related Structures:
4UX4, 4W5I - PubMed Abstract:
The tankyrases are members of the PARP superfamily; they poly(ADP-ribosyl)ate their target proteins using NAD(+) as a source of electrophilic ADP-ribosyl units. The three principal protein substrates of the tankyrases (TRF1, NuMA and axin) are involved in replication of cancer cells; thus inhibitors of the tankyrases may have anticancer activity. Using structure-based drug design and by analogy with known 3-arylisoquinolin-1-one and 2-arylquinazolin-4-one inhibitors, series of arylnaphthyridinones, arylpyridinopyrimidinones and their tetrahydro-derivatives were synthesised and evaluated in vitro. 7-Aryl-1,6-naphthyridin-5-ones, 3-aryl-2,6-naphthyridin-1-ones and 3-aryl-2,7-naphthyridin-1-ones were prepared by acid-catalysed cyclisation of the corresponding arylethynylpyridinenitriles or reaction of bromopyridinecarboxylic acids with β-diketones, followed by treatment with NH3. The 7-aryl-1,6-naphthyridin-5-ones were methylated at 1-N and reduced to 7-aryl-1-methyl-1,2,3,4-tetrahydro-1,6-naphthyridin-5-ones. Cu-catalysed reaction of benzamidines with bromopyridinecarboxylic acids furnished 2-arylpyrido[2,3-d]pyrimidin-4-ones. Condensation of benzamidines with methyl 1-benzyl-4-oxopiperidine-3-carboxylate and deprotection gave 2-aryl-5,6,7,8-tetrahydropyrido[4,3-d]pyrimidin-4-ones, aza analogues of the known inhibitor XAV939. Introduction of the ring-N in the arylnaphthyridinones and the arylpyridopyrimidinones caused >1000-fold loss in activity, compared with their carbocyclic isoquinolinone and quinazolinone analogues. However, the 7-aryl-1-methyl-1,2,3,4-tetrahydro-1,6-naphthyridin-5-ones showed excellent inhibition of the tankyrases, with some examples having IC50=2nM. One compound (7-(4-bromophenyl)-1-methyl-1,2,3,4-tetrahydro-1,6-naphthyridin-5-one) showed 70-fold selectivity for inhibition of tankyrase-2 versus tankyrase-1. The mode of binding was explored through crystal structures of inhibitors in complex with tankyrase-2.
Organizational Affiliation:
Medicinal Chemistry, Department of Pharmacy & Pharmacology, University of Bath, Claverton Down, Bath BA2 7AY, UK.