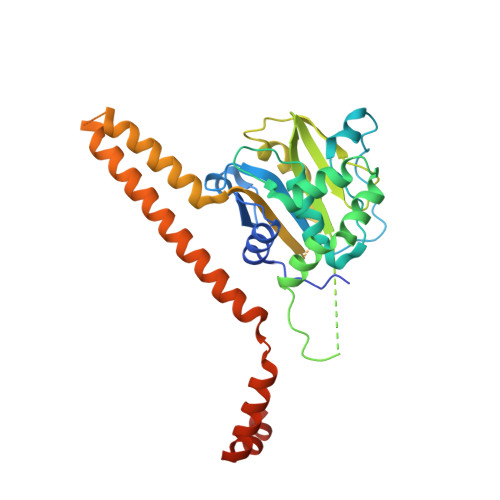
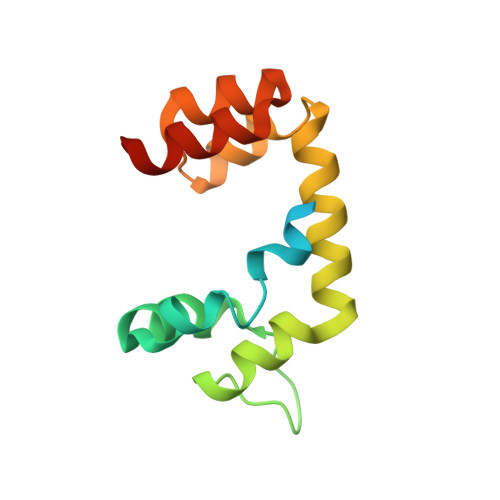
Mechanism of Uch-L5 Activation and Inhibition by Deubad Domains in Rpn13 and Ino80G.
Sahtoe, D.D., Van Dijk, W.J., El Oualid, F., Ekkebus, R., Ovaa, H., Sixma, T.K.(2015) Mol Cell 57: 887
- PubMed: 25702870
- DOI: https://doi.org/10.1016/j.molcel.2014.12.039
- Primary Citation of Related Structures:
4UEL, 4UEM, 4UF5, 4UF6 - PubMed Abstract:
Deubiquitinating enzymes (DUBs) control vital processes in eukaryotes by hydrolyzing ubiquitin adducts. Their activities are tightly regulated, but the mechanisms remain elusive. In particular, the DUB UCH-L5 can be either activated or inhibited by conserved regulatory proteins RPN13 and INO80G, respectively. Here we show how the DEUBAD domain in RPN13 activates UCH-L5 by positioning its C-terminal ULD domain and crossover loop to promote substrate binding and catalysis. The related DEUBAD domain in INO80G inhibits UCH-L5 by exploiting similar structural elements in UCH-L5 to promote a radically different conformation, and employs molecular mimicry to block ubiquitin docking. In this process, large conformational changes create small but highly specific interfaces that mediate activity modulation of UCH-L5 by altering the affinity for substrates. Our results establish how related domains can exploit enzyme conformational plasticity to allosterically regulate DUB activity. These allosteric sites may present novel insights for pharmaceutical intervention in DUB activity.
Organizational Affiliation:
Division of Biochemistry and Cancer Genomics Center, The Netherlands Cancer Institute, Plesmanlaan 121, 1066CX Amsterdam, the Netherlands.