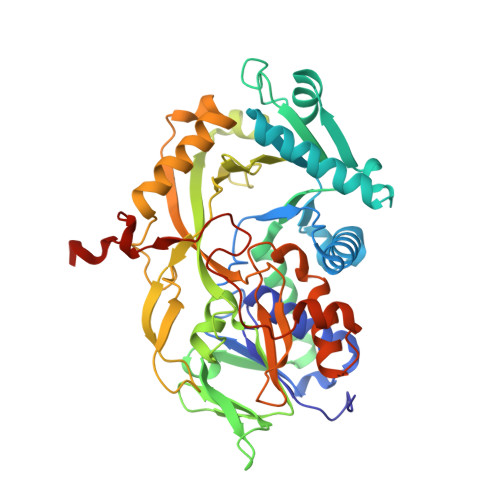
Structural basis of the substrate specificity of the FPOD/FAOD family revealed by fructosyl peptide oxidase from Eupenicillium terrenum
Gan, W., Gao, F., Xing, K., Jia, M., Liu, H., Gong, W.(2015) Acta Crystallogr F Struct Biol Commun 71: 381-387
- PubMed: 25849495
- DOI: https://doi.org/10.1107/S2053230X15003921
- Primary Citation of Related Structures:
4RSL - PubMed Abstract:
The FAOD/FPOD family of proteins has the potential to be useful for the longterm detection of blood glucose levels in diabetes patients. A bottleneck for this application is to find or engineer a FAOD/FPOD family enzyme that is specifically active towards α-fructosyl peptides but is inactive towards other types of glycated peptides. Here, the crystal structure of fructosyl peptide oxidase from Eupenicillium terrenum (EtFPOX) is reported at 1.9 Å resolution. In contrast to the previously reported structure of amadoriase II, EtFPOX has an open substrate entrance to accommodate the large peptide substrate. The functions of residues critical for substrate selection are discussed based on structure comparison and sequence alignment. This study reveals the first structural details of group I FPODs that prefer α-fructosyl substrates and could provide significant useful information for uncovering the mechanism of substrate specificity of FAOD/FPODs and guidance towards future enzyme engineering for diagnostic purposes.
Organizational Affiliation:
Key Laboratory of RNA, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Beijing 100101, People's Republic of China.