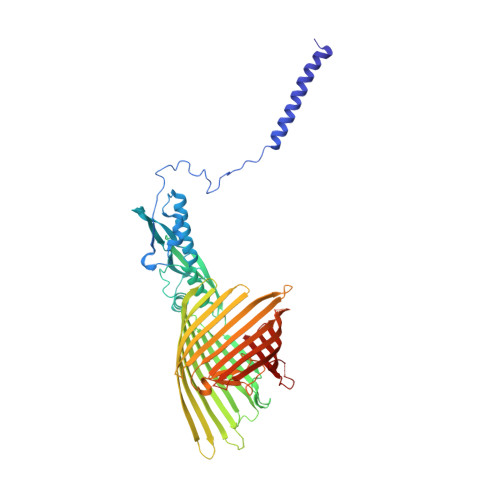
Conserved Omp85 lid-lock structure and substrate recognition in FhaC
Maier, T., Clantin, B., Gruss, F., Dewitte, F., Delattre, A.S., Jacob-Dubuisson, F., Hiller, S., Villeret, V.(2015) Nat Commun 6: 7452-7452
- PubMed: 26058369
- DOI: https://doi.org/10.1038/ncomms8452
- Primary Citation of Related Structures:
4QKY, 4QL0 - PubMed Abstract:
Omp85 proteins mediate translocation of polypeptide substrates across and into cellular membranes. They share a common architecture comprising substrate-interacting POTRA domains, a C-terminal 16-stranded β-barrel pore and two signature motifs located on the inner barrel wall and at the tip of the extended L6 loop. The observation of two distinct conformations of the L6 loop in the available Omp85 structures previously suggested a functional role of conformational changes in L6 in the Omp85 mechanism. Here we present a 2.5 Å resolution structure of a variant of the Omp85 secretion protein FhaC, in which the two signature motifs interact tightly and form the conserved 'lid lock'. Reanalysis of previous structural data shows that L6 adopts the same, conserved resting state position in all available Omp85 structures. The FhaC variant structure further reveals a competitive mechanism for the regulation of substrate binding mediated by the linker to the N-terminal plug helix H1.
Organizational Affiliation:
Biozentrum, University of Basel, Klingelbergstr. 70, Basel 4056, Switzerland.