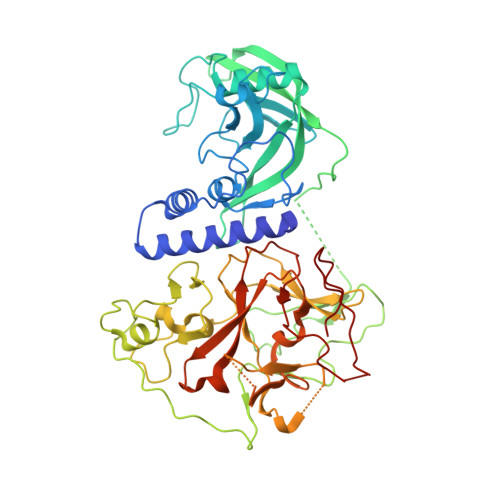
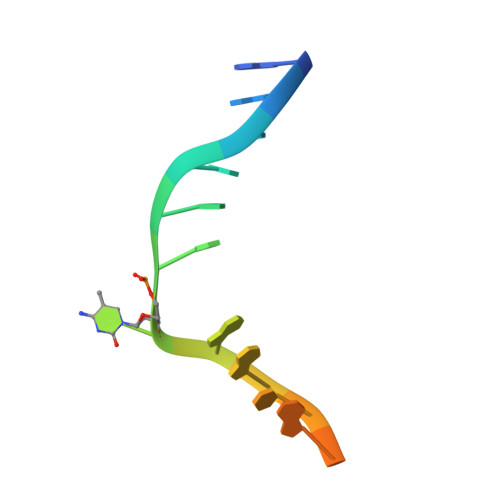
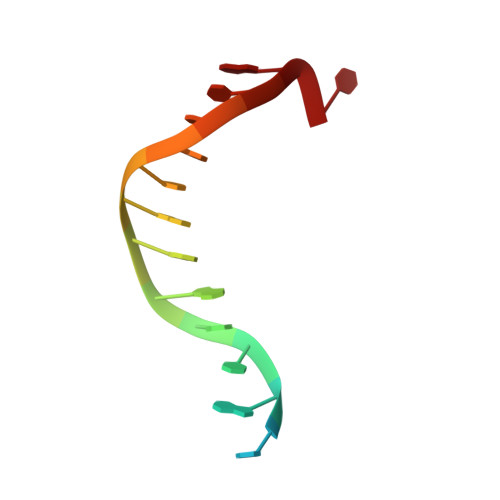
Mechanism of DNA Methylation-Directed Histone Methylation by KRYPTONITE.
Du, J., Johnson, L.M., Groth, M., Feng, S., Hale, C.J., Li, S., Vashisht, A.A., Gallego-Bartolome, J., Wohlschlegel, J.A., Patel, D.J., Jacobsen, S.E.(2014) Mol Cell 55: 495-504
- PubMed: 25018018
- DOI: https://doi.org/10.1016/j.molcel.2014.06.009
- Primary Citation of Related Structures:
4QEN, 4QEO, 4QEP - PubMed Abstract:
In Arabidopsis, CHG DNA methylation is controlled by the H3K9 methylation mark through a self-reinforcing loop between DNA methyltransferase CHROMOMETHYLASE3 (CMT3) and H3K9 histone methyltransferase KRYPTONITE/SUVH4 (KYP). We report on the structure of KYP in complex with methylated DNA, substrate H3 peptide, and cofactor SAH, thereby defining the spatial positioning of the SRA domain relative to the SET domain. The methylated DNA is bound by the SRA domain with the 5mC flipped out of the DNA, while the H3(1-15) peptide substrate binds between the SET and post-SET domains, with the ε-ammonium of K9 positioned adjacent to bound SAH. These structural insights, complemented by functional data on key mutants of residues lining the 5mC and H3K9-binding pockets within KYP, establish how methylated DNA recruits KYP to the histone substrate. Together, the structures of KYP and previously reported CMT3 complexes provide insights into molecular mechanisms linking DNA and histone methylation.
Organizational Affiliation:
Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10065, USA.