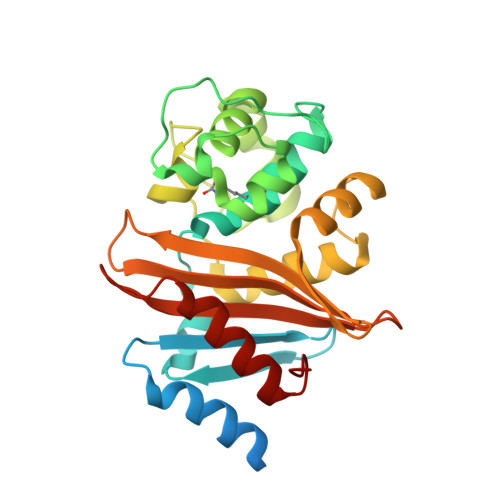
Crystal Structure of Carbapenemase OXA-58 from Acinetobacter baumannii.
Smith, C.A., Antunes, N.T., Toth, M., Vakulenko, S.B.(2014) Antimicrob Agents Chemother 58: 2135-2143
- PubMed: 24468777
- DOI: https://doi.org/10.1128/AAC.01983-13
- Primary Citation of Related Structures:
4OH0 - PubMed Abstract:
Class D β-lactamases capable of hydrolyzing last-resort carbapenem antibiotics represent a major challenge for treatment of bacterial infections. Wide dissemination of these enzymes in Acinetobacter baumannii elevated this pathogen to the category of most deadly and difficult to treat. We present here the structure of the OXA-58 β-lactamase, a major class D carbapenemase of A. baumannii, determined to 1.30-Å resolution. Unlike two other Acinetobacter carbapenemases, OXA23 and OXA-24, the OXA-58 enzyme lacks the characteristic hydrophobic bridge over the active site, despite conservation of the residues which participate in its formation. The active-site residues in OXA-58 are spatially conserved in comparison to those in other class D β-lactamases. Lys86, which activates water molecules during the acylation and deacylation steps, is fully carboxylated in the OXA-58 structure. In the absence of a substrate, a water molecule is observed in the active site of the enzyme and is positioned in the pocket that is usually occupied by the 6α-hydroxyethyl moiety of carbapenems. A water molecule in this location would efficiently deacylate good substrates, such as the penicillins, but in the case of carbapenems, it would be expelled by the 6α-hydroxyethyl moiety of the antibiotics and a water from the surrounding medium would find its way to the vicinity of the carboxylated Lys86 to perform deacylation. Subtle differences in the position of this water in the acyl-enzyme complexes of class D β-lactamases could ultimately be responsible for differences in the catalytic efficiencies of these enzymes against last-resort carbapenem antibiotics.
Organizational Affiliation:
Stanford Synchrotron Radiation Lightsource, Stanford University, Menlo Park, California, USA.