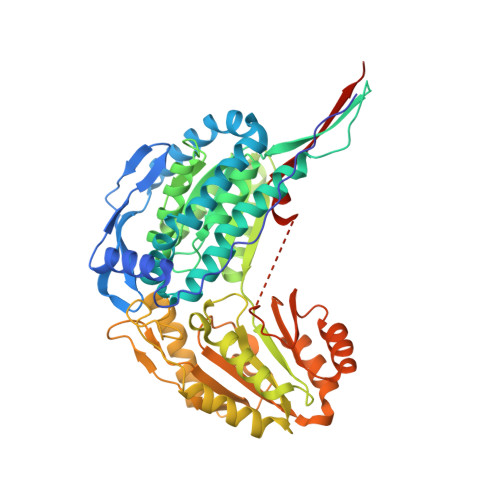
Structural Studies of Yeast Delta (1)-Pyrroline-5-carboxylate Dehydrogenase (ALDH4A1): Active Site Flexibility and Oligomeric State.
Pemberton, T.A., Srivastava, D., Sanyal, N., Henzl, M.T., Becker, D.F., Tanner, J.J.(2014) Biochemistry 53: 1350-1359
- PubMed: 24502590
- DOI: https://doi.org/10.1021/bi500048b
- Primary Citation of Related Structures:
4OE4, 4OE5, 4OE6 - PubMed Abstract:
The proline catabolic enzyme Δ(1)-pyrroline-5-carboxylate dehydrogenase (ALDH4A1) catalyzes the NAD(+)-dependent oxidation of γ-glutamate semialdehyde to l-glutamate. In Saccharomyces cerevisiae, ALDH4A1 is encoded by the PUT2 gene and known as Put2p. Here we report the steady-state kinetic parameters of the purified recombinant enzyme, two crystal structures of Put2p, and the determination of the oligomeric state and quaternary structure from small-angle X-ray scattering and sedimentation velocity. Using Δ(1)-pyrroline-5-carboxylate as the substrate, catalytic parameters kcat and Km were determined to be 1.5 s(-1) and 104 μM, respectively, with a catalytic efficiency of 14000 M(-1) s(-1). Although Put2p exhibits the expected aldehyde dehydrogenase superfamily fold, a large portion of the active site is disordered in the crystal structure. Electron density for the 23-residue aldehyde substrate-binding loop is absent, implying substantial conformational flexibility in solution. We furthermore report a new crystal form of human ALDH4A1 (42% identical to Put2p) that also shows disorder in this loop. The crystal structures provide evidence of multiple active site conformations in the substrate-free form of the enzyme, which is consistent with a conformational selection mechanism of substrate binding. We also show that Put2p forms a trimer-of-dimers hexamer in solution. This result is unexpected because human ALDH4A1 is dimeric, whereas some bacterial ALDH4A1s are hexameric. Thus, global sequence identity and domain of life are poor predictors of the oligomeric states of ALDH4A1. Mutation of a single Trp residue that forms knob-in-hole interactions across the dimer-dimer interface abrogates hexamer formation, suggesting that this residue is the center of a protein-protein association hot spot.
Organizational Affiliation:
Department of Chemistry, University of Missouri-Columbia , Columbia, Missouri 65211, United States.