Discovery of lead compounds targeting the bacterial sliding clamp using a fragment-based approach.
Yin, Z., Whittell, L.R., Wang, Y., Jergic, S., Liu, M., Harry, E.J., Dixon, N.E., Beck, J.L., Kelso, M.J., Oakley, A.J.(2014) J Med Chem 57: 2799-2806
- PubMed: 24592885
- DOI: https://doi.org/10.1021/jm500122r
- Primary Citation of Related Structures:
4N94, 4N95, 4N96, 4N97, 4N98, 4N99, 4N9A - PubMed Abstract:
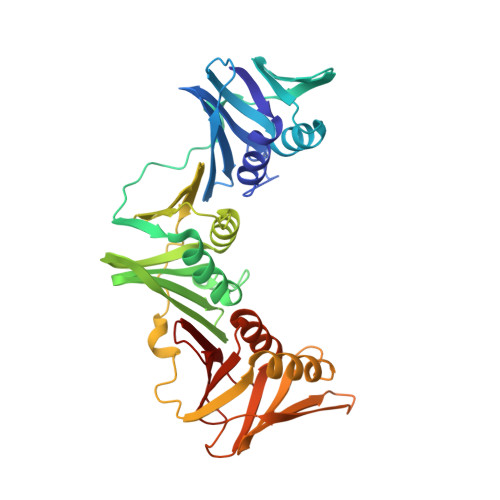
The bacterial sliding clamp (SC), also known as the DNA polymerase III β subunit, is an emerging antibacterial target that plays a central role in DNA replication, serving as a protein-protein interaction hub with a common binding pocket to recognize linear motifs in the partner proteins. Here, fragment-based screening using X-ray crystallography produced four hits bound in the linear-motif-binding pocket of the Escherichia coli SC. Compounds structurally related to the hits were identified that inhibited the E. coli SC and SC-mediated DNA replication in vitro. A tetrahydrocarbazole derivative emerged as a promising lead whose methyl and ethyl ester prodrug forms showed minimum inhibitory concentrations in the range of 21-43 μg/mL against representative Gram-negative and Gram-positive bacteria species. The work demonstrates the utility of a fragment-based approach for identifying bacterial sliding clamp inhibitors as lead compounds with broad-spectrum antibacterial activity.
Organizational Affiliation:
School of Chemistry and Centre for Medical and Molecular Bioscience, University of Wollongong , Wollongong, New South Wales 2522, Australia.