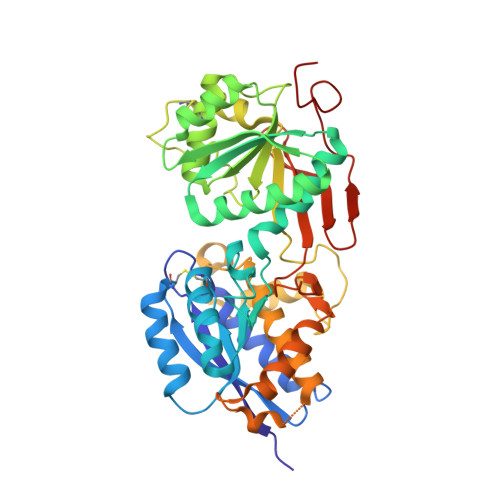
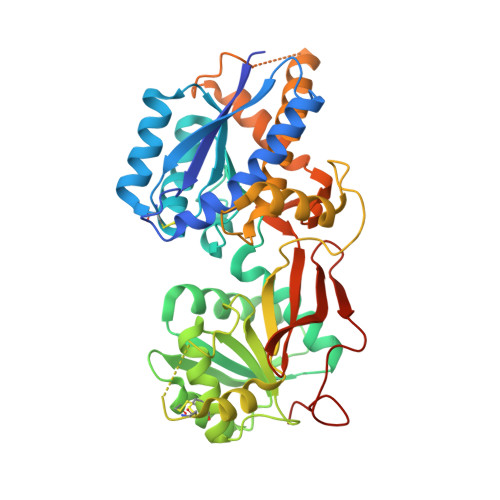
Structural mechanism of ligand activation in human GABA(B) receptor.
Geng, Y., Bush, M., Mosyak, L., Wang, F., Fan, Q.R.(2013) Nature 504: 254-259
- PubMed: 24305054
- DOI: https://doi.org/10.1038/nature12725
- Primary Citation of Related Structures:
4MQE, 4MQF, 4MR7, 4MR8, 4MR9, 4MRM, 4MS1, 4MS3, 4MS4 - PubMed Abstract:
Human GABA(B) (γ-aminobutyric acid class B) receptor is a G-protein-coupled receptor central to inhibitory neurotransmission in the brain. It functions as an obligatory heterodimer of the subunits GBR1 and GBR2. Here we present the crystal structures of a heterodimeric complex between the extracellular domains of GBR1 and GBR2 in the apo, agonist-bound and antagonist-bound forms. The apo and antagonist-bound structures represent the resting state of the receptor; the agonist-bound complex corresponds to the active state. Both subunits adopt an open conformation at rest, and only GBR1 closes on agonist-induced receptor activation. The agonists and antagonists are anchored in the interdomain crevice of GBR1 by an overlapping set of residues. An antagonist confines GBR1 to the open conformation of the inactive state, whereas an agonist induces its domain closure for activation. Our data reveal a unique activation mechanism for GABA(B) receptor that involves the formation of a novel heterodimer interface between subunits.
Organizational Affiliation:
Department of Pharmacology, Columbia University, New York, New York 10032, USA.