Elucidation of the crystal structure of Coriolopsis caperata laccase: restoration of the structure and activity of the native enzyme from the T2-depleted form by copper ions.
Glazunova, O.A., Polyakov, K.M., Fedorova, T.V., Dorovatovskii, P.V., Koroleva, O.V.(2015) Acta Crystallogr D Biol Crystallogr 71: 854-861
- PubMed: 25849396
- DOI: https://doi.org/10.1107/S1399004715001595
- Primary Citation of Related Structures:
4JHU, 4JHV - PubMed Abstract:
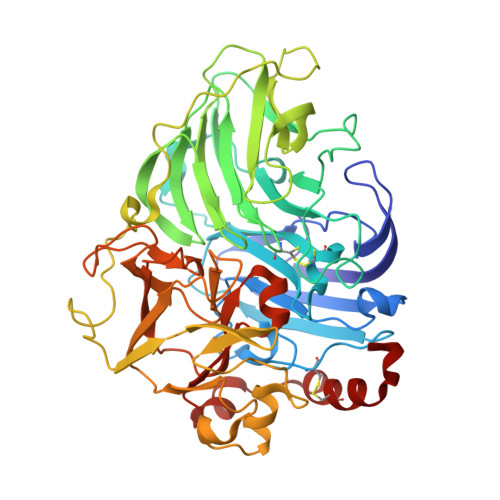
Laccases are members of a large family of multicopper oxidases that catalyze the oxidation of a wide range of organic and inorganic substrates accompanied by the reduction of dioxygen to water. A new laccase was isolated from the basidiomycete Coriolopsis caperata strain 0677 and its amino-acid sequence was determined. According to its physicochemical properties and spectroscopic features, the laccase from C. caperata is a high redox-potential blue laccase. Attempts to crystallize the native enzyme were unsuccessful. The copper type 2-depleted (T2D) laccase was prepared and crystallized. The structure of T2D laccase from C. caperata was solved at 1.6 Å resolution, and attempts to reconstruct the T2 copper centre were performed using Cu(+) and Cu(2+) ions. The structure of T2D+Cu(+) laccase was solved at 1.89 Å resolution. It was shown that the T2D+Cu(+) laccase structure contained four copper ions in the active site. Reconstruction could not be achieved when the T2D laccase crystals were treated with CuSO4.
Organizational Affiliation:
A. N. Bach Institute of Biochemistry, Russian Academy of Sciences, Leninsky Prospect 33, Moscow 119071, Russian Federation.