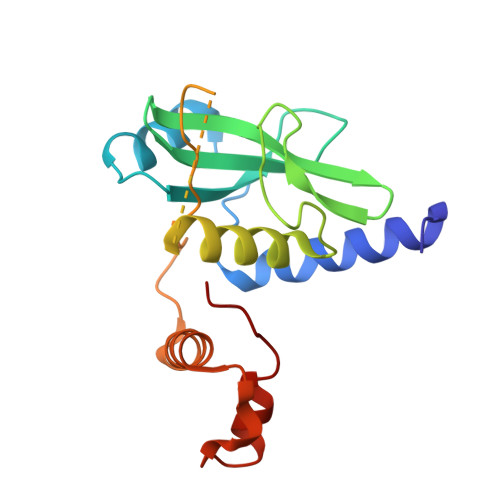
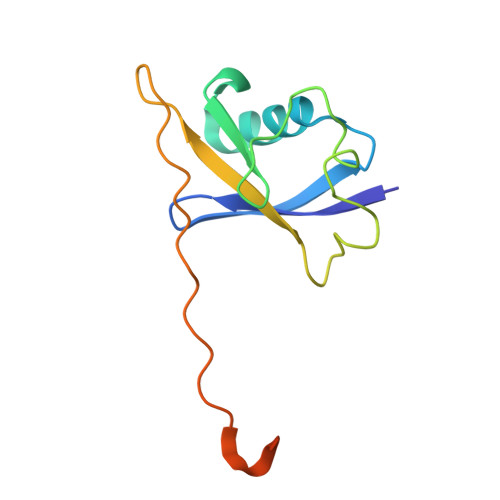
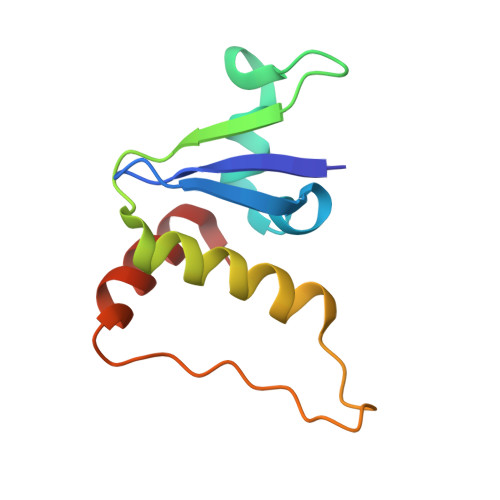
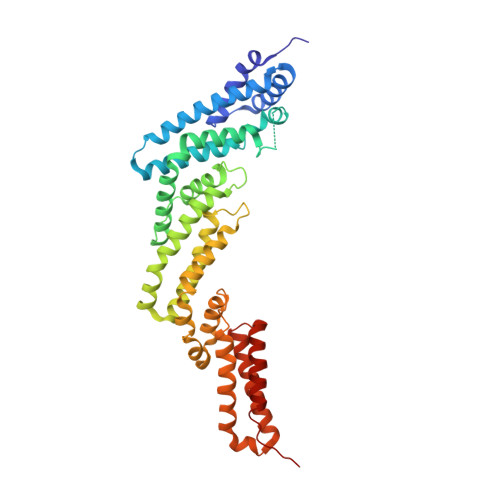
Structural basis of intersubunit recognition in elongin BC-cullin 5-SOCS box ubiquitin-protein ligase complexes.
Kim, Y.K., Kwak, M.J., Ku, B., Suh, H.Y., Joo, K., Lee, J., Jung, J.U., Oh, B.H.(2013) Acta Crystallogr D Biol Crystallogr 69: 1587-1597
- PubMed: 23897481
- DOI: https://doi.org/10.1107/S0907444913011220
- Primary Citation of Related Structures:
4JGH - PubMed Abstract:
The cullin-RING ubiquitin ligases are multisubunit complexes that ubiquitinate various proteins. Six different cullins encoded by the human genome selectively pair with different adaptors and substrate receptors. It is presently poorly understood how cullin-2 (Cul2) and cullin-5 (Cul5) associate specifically with their adaptor elongin BC and a SOCS-box-containing substrate receptor. Here, crystallographic and mutational analyses of a quaternary complex between the N-terminal half of Cul5, elongin BC and SOCS2 are reported. Cul5 interacts extensively with elongin BC via residues that are highly conserved in Cul2 but not in other cullins. Cul5 also interacts with SOCS2, but via only two residues, Pro184 and Arg186, which are located in the C-terminal part of the SOCS box called the Cul5 box. Pro184 makes a ring-to-ring interaction with Trp53 of Cul5, which is substituted by alanine in Cul2. This interaction is shown to contribute significantly to the overall binding affinity between Cul5 and SOCS2-elongin BC. This study provides structural bases underlying the specificity of Cul5 and Cul2 for elongin BC and their preferential association with Cul5 or Cul2 box-containing substrate receptors.
Organizational Affiliation:
Department of Biological Sciences, Korea Advanced Institute of Science and Technology, Daejeon 305-701, Republic of Korea.