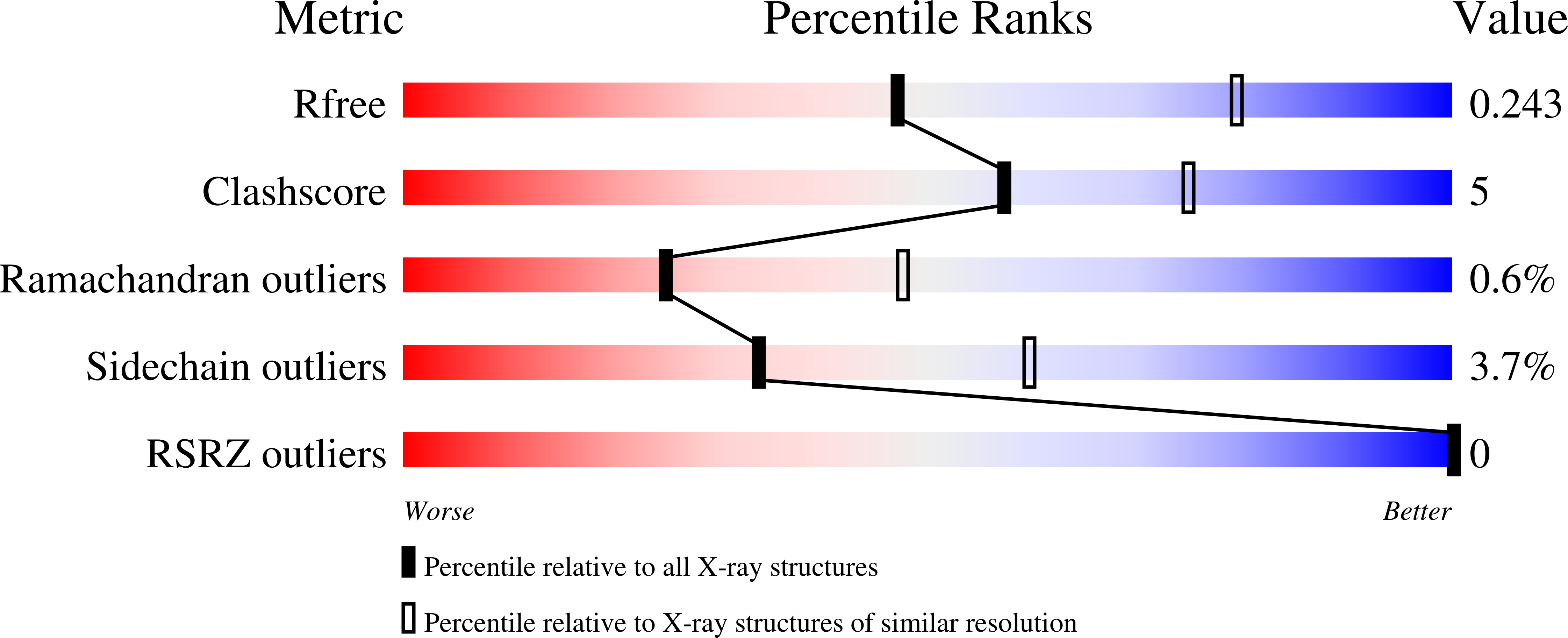
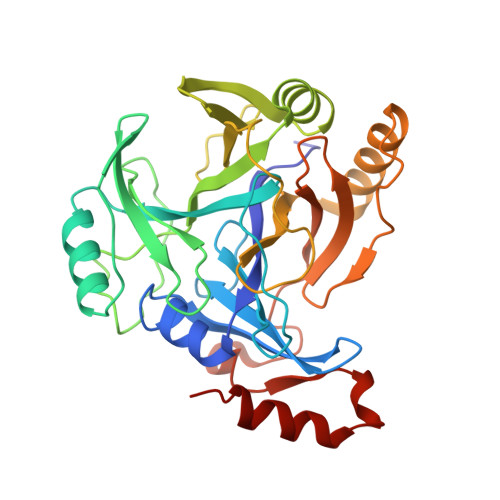
Structural characterization of 2,6-dichloro-p-hydroquinone 1,2-dioxygenase (PcpA) from Sphingobium chlorophenolicum, a new type of aromatic ring-cleavage enzyme.
Hayes, R.P., Green, A.R., Nissen, M.S., Lewis, K.M., Xun, L., Kang, C.(2013) Mol Microbiol 88: 523-536
- PubMed: 23489289
- DOI: https://doi.org/10.1111/mmi.12204
- Primary Citation of Related Structures:
4HUZ - PubMed Abstract:
PcpA (2,6-dichloro-p-hydroquinone 1,2-dioxygenase) from Sphingobium chlorophenolicum, a non-haem Fe(II) dioxygenase capable of cleaving the aromatic ring of p-hydroquinone and its substituted variants, is a member of the recently discovered p-hydroquinone 1,2-dioxygenases. Here we report the 2.6 Å structure of PcpA, which consists of four βαβββ motifs, a hallmark of the vicinal oxygen chelate superfamily. The secondary co-ordination sphere of the Fe(II) centre forms an extensive hydrogen-bonding network with three solvent exposed residues, linking the catalytic Fe(II) to solvent. A tight hydrophobic pocket provides p-hydroquinones access to the Fe(II) centre. The p-hydroxyl group is essential for the substrate-binding, thus phenols and catechols, lacking a p-hydroxyl group, do not bind to PcpA. Site-directed mutagenesis and kinetic analysis confirm the critical catalytic role played by the highly conserved His10, Thr13, His226 and Arg259. Based on these results, we propose a general reaction mechanism for p-hydroquinone 1,2-dioxygenases.
Organizational Affiliation:
Department of Chemistry, Washington State University, Pullman, WA 99164-4630, USA.